TruthArchive.ai - Tweets Saved By @DrHermiz

@DrHermiz - David J Hermiz MD
"You can manipulate [coronaviruses] in the lab pretty easily. Spike protein drives a lot of what happens with the coronavirus, zoonotic risk. So you can get the sequence, you can build the protein, and we work with Ralph Baric at UNC to do this. Insert it into the backbone of another virus and do some work in the labs." -- Dr. Peter Daszak, President of the EcoHealth Alliance American Society for Microbiology. This Week in Virology Podcast. (2020, May 19). Peter Daszak of EcoHealth Alliance - TWiV 615. https://asm.org/Podcasts/TWiV/Episodes/Peter-Daszak-of-EcoHealth-Alliance-TWiV-615


@DrHermiz - David J Hermiz MD
https://www.nature.com/articles/nm.3985 Researchers at UNC led by Dr. Ralph Baric in 2015 cultured SARS-CoV variants in specific cell lines to study infection patterns. This included using pseudoviruses (non-replicative, inert viruses also used in vaccine development) to assess how the virus might bind to human cells and developing growth curves to understand virus behavior in various environments. To create chimeric viruses, the researchers employed reverse genetics to manipulate plasmids—DNA molecules used to artificially introduce foreign DNA into a cell. This enabled insertion of the the spike protein gene into a viral genome and observing the resultant effects on host cell infection, virus replication, and pathogenesis in a controlled laboratory setting. This chimeric virus produced from the study was able effectively use the human ACE2 receptor, replicate in human airway cells, and cause disease in mice. Current SARS treatments were ineffective against it, indicating a risk of new SARS-CoV outbreaks from bat populations.
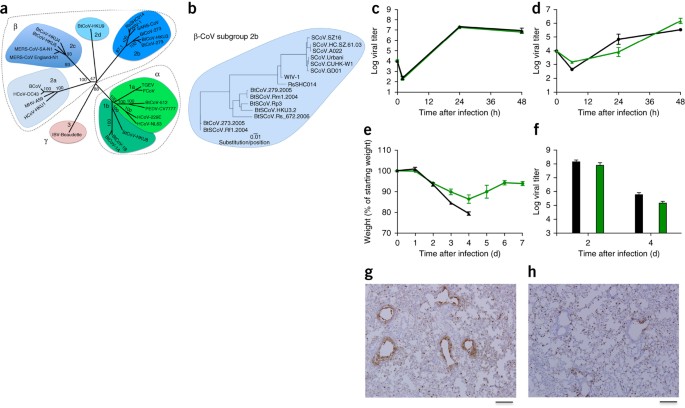

@DrHermiz - David J Hermiz MD
Dr. Peter Daszak has been ordered to testify before the US Congress in Washington D.C. on November 14, 2023, regarding his alleged involvement in the creation and accidental release of SARS-CoV-2, and his involvement in a conspiracy to cover this up. #OriginOfCovid #COVID19

@DrHermiz - David J Hermiz MD
Dr. Peter Daszak, as the president of the @EcoHealthNYC, submitted a funding request to the Defense Advanced Research Projects Agency (@DARPA), an agency of the United States Department of Defense (@DeptofDefense) in 2018. The title of the project was "Defusing the Threat of Bat-borne Coronaviruses." The project entailed the insertion of a furin cleavage site into SARS-like coronaviruses gathered from their natural reservoir, located 1500 km away in southern China, to the research being conducted at the Wuhan Institute of Virology. The study of the furin protease cleavage site within the spike protein of SARS-CoV-2 remains an active area of research because of its critical function in how the virus gains entry into human cells. The "furin cleavage site" refers to a segment of the spike protein in a coronavirus that interacts with the human furin enzyme, a protease that catalyzes the reaction of cleaving the spike protein, thus activating the virus for entry into the human cell. @PeterDaszak has been served a subpoena by @COVIDSelect to determine his role in the probable accidental release of #SARSCoV2 from the Wuhan Institute of Virology. He has been ordered to testify in Washington D.C. on November 14, 2023, regarding his alleged involvement in the creation and accidental release of SARS-CoV-2, the causative agent of #COVID19, and his involvement in a conspiracy to cover this up. https://www.documentcloud.org/documents/21066966-defuse-proposal


@DrHermiz - David J Hermiz MD
So @Bryce_Nickels, a Professor of Genetics at Rutgers who is published in Science, Cell, Nature, and Nature Communications, gets blocked by the Editor-in-Chief of @NatureMedicine because he has been asking for a formal response to a petition signed by 5000 scientists and policymakers regarding his concerns about #ProximalOrigins, the seminal paper on the #OriginOfCovid. A congressional investigative committee already concluded, "Many of the conclusions reached by 'Proximal Origin' have never been proven or verified....Dr. Andersen made it clear that the co-authors allowed politics to influence the science behind “Proximal Origin."' When are we going to say enough is enough with people like @JMinImmunoland in these positions when they lack accountability and ethics, using their positions of influence to push their personal political agendas? How did we let these people attain these positions? https://oversight.house.gov/release/wenstrup-releases-alarming-new-report-on-proximal-origin-authors-nih-suppression-of-the-covid-19-lab-leak-hypothesis/


@DrHermiz - David J Hermiz MD
So @Bryce_Nickels, a Professor of Genetics at Rutgers who is published in Science, Cell, Nature, and Nature Communications, gets blocked by the Editor-in-Chief of @NatureMedicine because he has been asking for a formal response to a petition signed by dozens of scientists and policymakers regarding his concerns about #ProximalOrigins, the seminal paper on the #OriginOfCovid. A congressional investigative committee already concluded, "Many of the conclusions reached by 'Proximal Origin' have never been proven or verified....Dr. Andersen made it clear that the co-authors allowed politics to influence the science behind “Proximal Origin."' When are we going to say enough is enough with people like @JMinImmunoland in these positions when they lack accountability and ethics, using their positions of influence to push their personal political agendas? How did we let these people attain these positions? https://oversight.house.gov/release/wenstrup-releases-alarming-new-report-on-proximal-origin-authors-nih-suppression-of-the-covid-19-lab-leak-hypothesis/


@DrHermiz - David J Hermiz MD
"When you get a sequence of a virus and it looks like a relative of a known nasty pathogen, just like we did with SARS, we find other coronaviruses in bats, a whole house of them. Some of them looked very similar to SARS, so we sequenced the Spike protein—the protein that attaches to cells—then we—well, I didn't do this work, but my colleagues in China did the work—you create pseudo-particles, you insert the spike proteins from those, sift and bind to human cells. Each step of this brings you closer and closer to understanding how this virus could really become pathogenic in people." Dr. Peter Daszak, President of the EcoHealth Alliance, March 26, 2016, New York City Academy of Medicine, New York City, New York, USA, Pandemics Forum. #OriginOfCovid