reSee.it - Tweets Saved By @P_J_Buckhaults

@P_J_Buckhaults - Phillip Buckhaults
A PCR protocol for monitoring the purity of covid19 mRNA vaccines and their recipients. share with whomever might find such a protocol useful. Step 1. Order this gBLOCK synthetic positive control from IDTDNA dot COM CTACGTGAACCATCACCCTAATCAAGTTTTTTGAGGTCGAGGTGCCGTAAAGCACTAAATCGGAACCCTAAAGGGAGCCCCCGATTTAGAGATTGACGGGGAAAGCCGGCGAACGTGGCGAGAAAGGAAGGGAAGAAAGCGtccatctgaagcgagtcgctTGTGGAATGTGTGTCAGTTAGGGTGTGGAAAGTACCCAGGCTCCCCAGCAGGCAGAAGTATGCAAAGCATGCAACTCAATTAGTCAGCAACCAGGTGTGttcggaagatgccagtgtgaCGTTGGCTACCCGTGATATTGCTGAAGAACTTGACGGCGAATGGGCTGACCGCTTCCTCGTGCTTTACGGTATCGCCGCTCCCGATTCACAGCGCATCGCCTTCTATCGCCTTCTTGACGAGaatggacagcgtttgcttaaCTTGGGTGGAGAGGCTATTCGGCTATGACTGGGCACAACAGACAATCGGCTGCTCTGATGCCGCCGTGTTCCGACTGTCAGCGCAGGGGCGCCCGGTTCTTTTTGTCAAGACCGACCTGTCCGGTGACCTGAATGAACTGCAAGACGAGGgccccaaagccctcgtagacCTACAGCGTGAGCTATGAGAAAGCGCCACACTTCCCGAAGGGAGAAAGGCGGACAGGTATCCGGTAAGCGGCAGGGTCGGAACAGGAGAGCGCACAAGGGAGCTTCCAGGGGGAAACGCCTGGTATCTTTATAGTCCTGTCGGGTTTCGagcatttcacgcgactagccGTTTGTTTGCCGGATCAAGAGCTACCAAATCTTTTTCCGAAGGTAACTGGCTTCAGCAGAGCGCAGATAACAAATACTGTTCTTCTAGTGTAGCCGTAGTTAGtagatctcttcctactagtcCAGCTACCAGACACAGACAAACAGCCCTCGGAGAGACAGAAGCGTGGCCAGCCAGAGCATCATTGCCTACACAATGTCTCTGGGCACCGAGAACAGCGTGGCCTACTCCAACAACTCTATCGCaaaatggcattagcgattgtACATCTGGCTGGGCTTTATCGCCGGACTGATTGCCATCGTGATGATCACAATCATGCTGTGTTGCATGACCAGCTGCTGTAGCTGCCTGAAGGGCTGTTGTAGCTGTGACAGCTGCTGCAAGTTCGACGAGGACGATTacaatcgtacgaaatggaggAGCCTACACCAACAGCTTTACCAGAGGCATGTACTACCCCGACAAGGTGTTCAGATCCAGCGTGCTGCACTATACCCAGGACCTGTTCCTGCCTTTCTTCAgcactctaatgcaaatgcagGATCGCCGACTACAACTACAAGCTGCCCGACGACTTCACAGGCTGTGTGATTGCCTGGAACAGCAACAACCTGGACTCCAAAGTCGACGGCAACTACAATTACCTGTACCGGCTGTTCctgacttctgatcaacctcgCTACATACCTCGCTCTGCTAATCCTGTTACCAGTAGCTGCTGCCAGTGGCGATAAGTCGTGTCTTACCGGGTTAGACTCAAGACGATAGTTACCGGATAAGGCGCtttccgtcccacctggtttaAGATGGCCTACCGGTTCAACGGCATCGGAGTGACACAGAATGTGCTGTACGAGAACCAGAAGCTGATCGCCAACCAGTTCAACAACGCCATCGGCAAGATCCAGGACAGCCTGA
@P_J_Buckhaults - Phillip Buckhaults
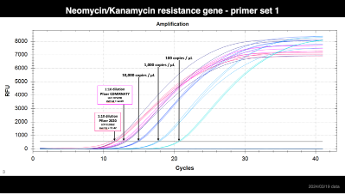
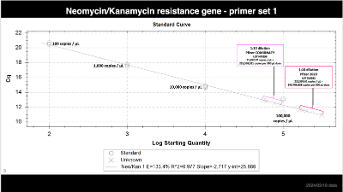
Step 1. This is a picture of the gblock standard and the locations of the primers used in the PCR. https://t.co/og2em7vmMO
@P_J_Buckhaults - Phillip Buckhaults
Step 2. Prepare serial dilutions of this gBLOCK standard. We use these concentrations and we dilute them in a solution of 0.3ng/ul human genomic DNA.. the HGDNA just prevents the gBLOCK standard from absorbing to the walls of the tube. Skip the HGDNA if you want to make fresh standard every week or if you use low-bind tubes. a. 100,000 copies / 2ul b. 10,000 copies / 2ul c. 1,000 copies / 2ul d. 100 copies / 2ul e. 10 copies / 2ull f. 0 copies / 2ul
@P_J_Buckhaults - Phillip Buckhaults
Step 3.Order these PCR primers and TaqMan probes from IDTDNA dot COM SV40 FWDTGTGGAATGTGTGTCAGTTAGG SV40 REVCACACCTGGTTGCTGACTAAT SV40 PRB/56-FAM/CCAGCAGGC/ZEN/AGAAGTATGCAAAGC/3IABkFQ/ Neo1 FWDCGTTGGCTACCCGTGATATT Neo1 REVCTCGTCAAGAAGGCGATAGAAG Neo1 PRB/56-FAM/CCGCTTCCT/ZEN/CGTGCTTTACGGTAT/3IABkFQ/ Neo5 FWDACTGGGCACAACAGACAATC Neo5REVCCTCGTCTTGCAGTTCATTCA Neo5 PRB/56-FAM/TTTGTCAAG/ZEN/ACCGACCTGTCCGG/3IABkFQ/ pBR322-5 FWDGTTTGTTTGCCGGATCAAGAG pBR322-5 REVCTAACTACGGCTACACTAGAAGAAC pBR322-5 PRB/56-FAM/TTCCGAAGG/ZEN/TAACTGGCTTCAGCA/3IABkFQ/ Spike1 FWDCAGCTACCAGACACAGACAAA Spike1 REVGCGATAGAGTTGTTGGAGTAGG Spike1 PRB/56-FAM/AGCCAGAGC/ZEN/ATCATTGCCTACACA/3IABkFQ/ Spike N1 FWDAGCCTACACCAACAGCTTTAC Spike N1 REVTGAAGAAAGGCAGGAACAGG Spike N1 PRB/56-FAM/CGACAAGGT/ZEN/GTTCAGATCCAGCGT/3IABkFQ/
@P_J_Buckhaults - Phillip Buckhaults
Step 4. Prepare these PCR primer / probe mixes. Do this for each of the above forward primer/reverse primer / probe combinations. You will end up with six different F/R/P mixes that detect different locations around the plasmid. Prepare 500ul of a 10X Primer/Probe Mix H2O437.5ul Forward primer [100uM]25.0ul Reverse primer [100uM]25.0ul PrimeTime Probe [100uM]12.5ul TOTAL Volume500ul
@P_J_Buckhaults - Phillip Buckhaults
Step 5. Make your PCR master mix. This is the recipe for the PCR reaction. It works on raw vaccine, raw biological fluids (diluted 10 fold in water) or tumor DNA (fresh frozen or FFPE). For a single 20ul PCR reaction…. H2O6ul 2x Quantabio PerfecTCa ToughMix UNG (95138-012)10ul 10x Primer/Probe Mix [5μMF, 5μMR, 2.5μMProbe]2ul DNA Template (straight vaccine diluted 10-fold in water works)2ul TOTAL20ul We typically make a master mix sufficient for all the samples, negative controls and quantification standards and aliquot 18ul/well into the PCR plate, then add the template (2ul) to the appropriate wells.
@P_J_Buckhaults - Phillip Buckhaults
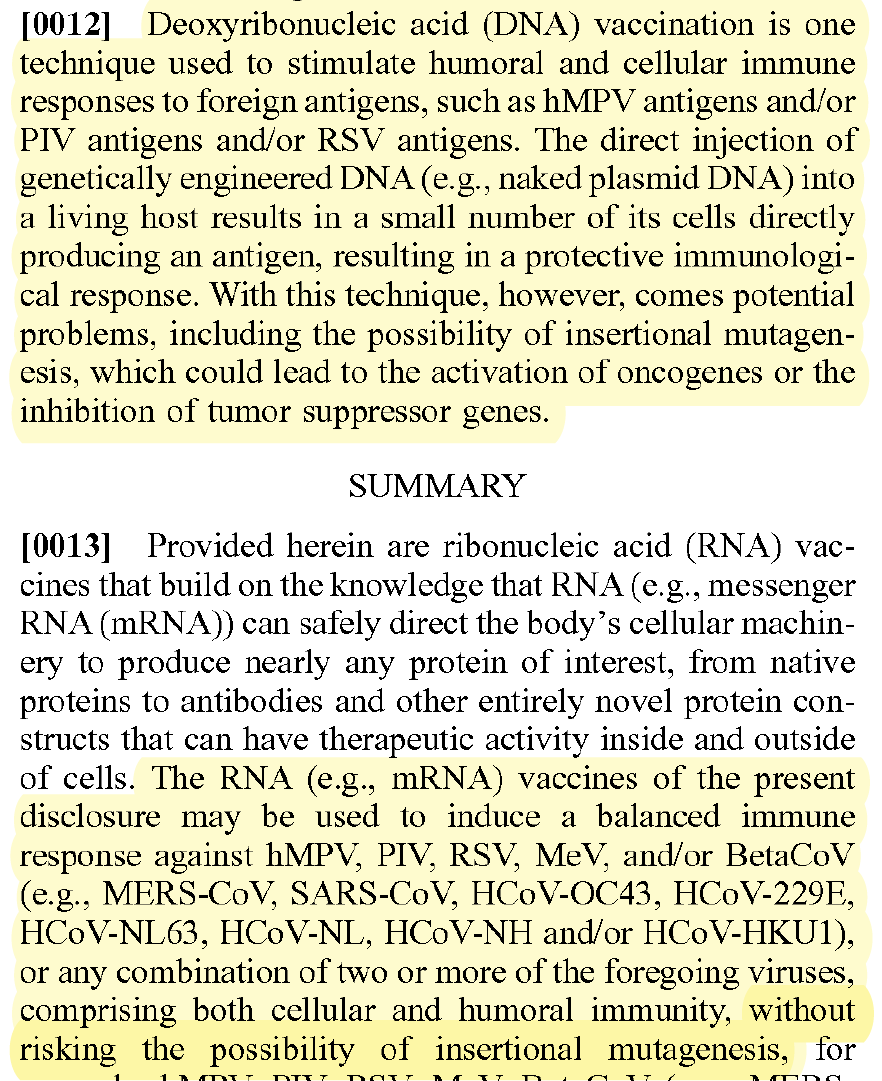
Step 6. Touchdown PCR cycling protocol for a BioRad CFX Opus 96 realtime PCR system. 1: 45.0°C for 5:00 (this is the uracil n-glycosylase incubation step to remove carryover contaminants. Only needed for UNG reagents) 2: 95.0°C for 2:00 3: 95.0°C for 0:10 4: 65.0°C for 0:10 5: GOTO 3, 3 more times 6: 95.0°C for 0:10 7: 62.0°C for 0:10 8: GOTO 6, 3 more times 9: 95.0°C for 0:10 10: 59.0°C for 0:10 11: GOTO 9, 3 more times 12: 95.0°C for 0:10 13: 58.0°C for 0:10 Plate Read 14: GOTO 12, 40 more times
@P_J_Buckhaults - Phillip Buckhaults
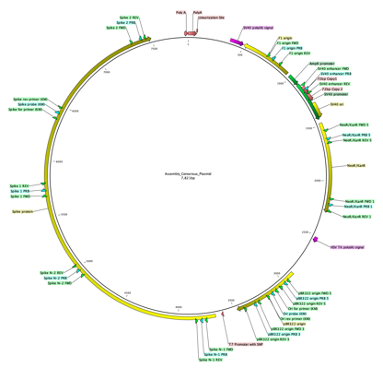
A circular map of the plasmid used in the manufacture of one of the mRNA vaccines. https://t.co/xnXCm07XAz
@P_J_Buckhaults - Phillip Buckhaults
The fastA sequence of what was retrieved from two batches of vaccine by nanopore sequencing. This sequence was used in the primer design and can be used as a reference sequence for the assembly of existing Illumina or Nanopore genome sequencing reads. >Assembly_Consensus_Plasmid AAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAA AAAAAAAAAAAAAAAAGAAGAGCTCCAACCGGTGTGGTAGCTCCGCCGTTTAACATCGCC CTTCCCAACAGTTGCGCAGCCTGAATGGCGAATGGAGATCCAATTTTTAAGTGTATAATG TGTTAAACTACTGATTCTAATTGTTTGTGTATTTTAGATTCACAGTCCCAAGGCTCATTT CAGGCCCCTCAGTCCTCACAGTCTGTTCATGATCATAATCAGCCATACCACATTTGTAGA GGTTTTACTTGCTTTAAAAAACCTCCCACACCTCCCCCTGAACCTGAAACATAAAATGAA TGCAATTGTTGTTGTTAACTTGTTTATTGCAGCTTATAATGGTTACAAATAAAGCAATAG CATCACAAATTTCACAAATAAAGCATTTTTTTCACTGCATTCTAGTTGTGGTTTGTCCAA ACTCATCAATGTATCTTAACGCGTAAATTGTAAGCGTTAATATTTTGTTAAAATTCGCGT TAAATTTTTGTTAAATCAGCTCATTTTTTAACCAATAGGCCGAAATCGGCAAAATCCCTT ATAAATCAAAAGAATAGACCGAGATAGGGTTGAGTGTTGTTCCAGTTTGGAACAAGAGTC CACTATTAAAGAACGTGGACTCCAACGTCAAAGGGCGAAAAACCGTCTATCAGGGCGATG GCCCACTACGTGAACCATCACCCTAATCAAGTTTTTTGGGGTCGAGGTGCCGTAAAGCAC TAAATCGGAACCCTAAAGGGAGCCCCCGATTTAGAGCTTGACGGGGAAAGCCGGCGAACG TGGCGAGAAAGGAAGGGAAGAAAGCGAAAGGAGCGGGCGCTAGGGCGCTGGCAAGTGTAG CGGTCACGCTGCGCGTAACCACCACACCCGCCGCGCTTAATGCGCCGCTACAGGGCGCGT CAGGTGGCACTTTTCGGGGAAATGTGCGCGGAACCCCTATTTGTTTATTTTTCTAAATAC ATTCAAATATGTATCCGCTCATGAGACAATAACCCTGATAAATGCTTCAATAATATTGAA AAAGGAAGAATCCTGAGGCGGAAAGAACCAGCTGTGGAATGTGTGTCAGTTAGGGTGTGG AAAGTCCCCAGGCTCCCCAGCAGGCAGAAGTATGCAAAGCATGCATCTCAATTAGTCAGC AACCAGGTGTGGAAAGTCCCCAGGCTCCCCAGCAGGCAGAAGTATGCAAAGCATGCATCT CAATTAGTCAGCAACCATAGTCCCGCCCCTAACTCCGCCCATCCCGCCCCTAACTCCGCC CAGTTCCGCCCATTCTCCGCCCCATGGCTGACTAATTTTTTTTATTTATGCAGAGGCCGA GGCCGCCTCGGCCTCTGAGCTATTCCAGAAGTAGTGAGGAGGCTTTTTTGGAGGCCTAGG CTTTTGCAAAGATCGATCAAGAGACAGGATGAGGATCGTTTCGCATGATTGAACAAGATG GATTGCACGCAGGTTCTCCGGCCGCTTGGGTGGAGAGGCTATTCGGCTATGACTGGGCAC AACAGACAATCGGCTGCTCTGATGCCGCCGTGTTCCGGCTGTCAGCGCAGGGGCGCCCGG TTCTTTTTGTCAAGACCGACCTGTCCGGTGCCCTGAATGAACTGCAAGACGAGGCAGCGC GGCTATCGTGGCTGGCCACGACGGGCGTTCCTTGCGCAGCTGTGCTCGACGTTGTCACTG AAGCGGGAAGGGACTGGCTGCTATTGGGCGAAGTGCCGGGGCAGGATCTCCTGTCATCTC ACCTTGCTCCTGCCGAGAAAGTATCCATCATGGCTGATGCAATGCGGCGGCTGCATACGC TTGATCCGGCTACCTGCCCATTCGACCACCAAGCGAAACATCGCATCGAGCGAGCACGTA CTCGGATGGAAGCCGGTCTTGTCGATCAGGATGATCTGGACGAAGAACATCAGGGGCTCG CGCCAGCCGAACTGTTCGCCAGGCTCAAGGCGAGCATGCCCGACGGCGAGGATCTCGTCG TGACCCATGGCGATGCCTGCTTGCCGAATATCATGGTGGAAAATGGCCGCTTTTCTGGAT TCATCGACTGTGGCCGGCTGGGTGTGGCGGACCGCTATCAGGACATAGCGTTGGCTACCC GTGATATTGCTGAAGAACTTGGCGGCGAATGGGCTGACCGCTTCCTCGTGCTTTACGGTA TCGCCGCTCCCGATTCGCAGCGCATCGCCTTCTATCGCCTTCTTGACGAGTTCTTCTGAG CGGGACTCTGGGGTTCGAAATGACCGACCAAGCGACGCCCAACCTGCCATCACGAGATTT CGATTCCACCGCCGCCTTCTATGAAAGGTTGGGCTTCGGAATCGTTTTCCGGGACGCCGG CTGGATGATCCTCCAGCGCGGGGATCTCATGCTGGAGTTCTTCGCCCACCCTAGGGGGAG GCTAACTGAAACACGGAAGGAGACAATACCGGAAGGAACCCGCGCTATGACGGCAATAAA AAGACAGAATAAAACGCACGGTGTTGGGTCGTTTGTTCATAAACGCGGGGTTCGGTCCCA GGGCTGGCACTCTGTCGATACCCCACCGAGACCCCATTGGGGCCAATACGCCCGCGTTTC TTCCTTTTCCCCACCCCACCCCCCAAGTTCGGGTGAAGGCCCAGGGCTCGCAGCCAACGT CGGGGCGGCAGGCCCTGCCATAGCCTCAGGTTACTCATATATACTTTAGATTGATTTAAA ACTTCATTTTTAATTTAAAAGGATCTAGGTGAAGATCCTTTTTGATAATCTCATGACCAA AATCCCTTAACGTGAGTTTTCGTTCCACTGAGCGTCAGACCCCGTAGAAAAGATCAAAGG ATCTTCTTGAGATCCTTTTTTCTGCGCGTAATCTGCTGCTTGCAAACAAAAAAACCACCG CTACCAGCGGTGGTTTGTTTGCCGGATCAAGAGCTACCAACTCTTTTTCCGAAGGTAACT GGCTTCAGCAGAGCGCAGATACCAAATACTGTTCTTCTAGTGTAGCCGTAGTTAGGCCAC CACTTCAAGAACTCTGTAGCACCGCCTACATACCTCGCTCTGCTAATCCTGTTACCAGTG GCTGCTGCCAGTGGCGATAAGTCGTGTCTTACCGGGTTGGACTCAAGACGATAGTTACCG GATAAGGCGCAGCGGTCGGGCTGAACGGGGGGTTCGTGCACACAGCCCAGCTTGGAGCGA ACGACCTACACCGAACTGAGATACCTACAGCGTGAGCTATGAGAAAGCGCCACGCTTCCC GAAGGGAGAAAGGCGGACAGGTATCCGGTAAGCGGCAGGGTCGGAACAGGAGAGCGCACG AGGGAGCTTCCAGGGGGAAACGCCTGGTATCTTTATAGTCCTGTCGGGTTTCGCCACCTC TGACTTGAGCGTCGATTTTTGTGATGCTCGTCAGGGGGGCGGAGCCTATGGAAAAACGCC AGCAACGCGGCCTTTTTACGGTTCCTGGCCTTTTGCTGGCCTTTTGCTCACATGTTCTTT CCTGCGTTATCCCCTGATTCTGTGGATAACCGTATTACCGCCATGCATTAGTTATTAATT AATACGACTCACTATAAGAATAAACTAGTATTCTTCTGGTCCCCACAGACTCAGAGAGAA CCCGCCACCATGTTCGTGTTCCTGGTGCTGCTGCCTCTGGTGTCCAGCCAGTGTGTGAAC CTGACCACCAGAACACAGCTGCCTCCAGCCTACACCAACAGCTTTACCAGAGGCGTGTAC TACCCCGACAAGGTGTTCAGATCCAGCGTGCTGCACTCTACCCAGGACCTGTTCCTGCCT TTCTTCAGCAACGTGACCTGGTTCCACGCCATCCACGTGTCCGGCACCAATGGCACCAAG AGATTCGACAACCCCGTGCTGCCCTTCAACGACGGGGTGTACTTTGCCAGCACCGAGAAG TCCAACATCATCAGAGGCTGGATCTTCGGCACCACACTGGACAGCAAGACCCAGAGCCTG CTGATCGTGAACAACGCCACCAACGTGGTCATCAAAGTGTGCGAGTTCCAGTTCTGCAAC GACCCCTTCCTGGGCGTCTACTACCACAAGAACAACAAGAGCTGGATGGAAAGCGAGTTC CGGGTGTACAGCAGCGCCAACAACTGCACCTTCGAGTACGTGTCCCAGCCTTTCCTGATG GACCTGGAAGGCAAGCAGGGCAACTTCAAGAACCTGCGCGAGTTCGTGTTTAAGAACATC GACGGCTACTTCAAGATCTACAGCAAGCACACCCCTATCAACCTCGTGCGGGATCTGCCT CAGGGCTTCTCTGCTCTGGAACCCCTGGTGGATCTGCCCATCGGCATCAACATCACCCGG TTTCAGACACTGCTGGCCCTGCACAGAAGCTACCTGACACCTGGCGATAGCAGCAGCGGA TGGACAGCTGGTGCCGCCGCTTACTATGTGGGCTACCTGCAGCCTAGAACCTTCCTGCTG AAGTACAACGAGAACGGCACCATCACCGACGCCGTGGATTGTGCTCTGGATCCTCTGAGC GAGACAAAGTGCACCCTGAAGTCCTTCACCGTGGAAAAGGGCATCTACCAGACCAGCAAC TTCCGGGTGCAGCCCACCGAATCCATCGTGCGGTTCCCCAATATCACCAATCTGTGCCCC TTCGGCGAGGTGTTCAATGCCACCAGATTCGCCTCTGTGTACGCCTGGAACCGGAAGCGG ATCAGCAATTGCGTGGCCGACTACTCCGTGCTGTACAACTCCGCCAGCTTCAGCACCTTC AAGTGCTACGGCGTGTCCCCTACCAAGCTGAACGACCTGTGCTTCACAAACGTGTACGCC GACAGCTTCGTGATCCGGGGAGATGAAGTGCGGCAGATTGCCCCTGGACAGACAGGCAAG ATCGCCGACTACAACTACAAGCTGCCCGACGACTTCACCGGCTGTGTGATTGCCTGGAAC AGCAACAACCTGGACTCCAAAGTCGGCGGCAACTACAATTACCTGTACCGGCTGTTCCGG AAGTCCAATCTGAAGCCCTTCGAGCGGGACATCTCCACCGAGATCTATCAGGCCGGCAGC ACCCCTTGTAACGGCGTGGAAGGCTTCAACTGCTACTTCCCACTGCAGTCCTACGGCTTT CAGCCCACAAATGGCGTGGGCTATCAGCCCTACAGAGTGGTGGTGCTGAGCTTCGAACTG CTGCATGCCCCTGCCACAGTGTGCGGCCCTAAGAAAAGCACCAATCTCGTGAAGAACAAA TGCGTGAACTTCAACTTCAACGGCCTGACCGGCACCGGCGTGCTGACAGAGAGCAACAAG AAGTTCCTGCCATTCCAGCAGTTTGGCCGGGATATCGCCGATACCACAGACGCCGTTAGA GATCCCCAGACACTGGAAATCCTGGACATCACCCCTTGCAGCTTCGGCGGAGTGTCTGTG ATCACCCCTGGCACCAACACCAGCAATCAGGTGGCAGTGCTGTACCAGGACGTGAACTGT ACCGAAGTGCCCGTGGCCATTCACGCCGATCAGCTGACACCTACATGGCGGGTGTACTCC ACCGGCAGCAATGTGTTTCAGACCAGAGCCGGCTGTCTGATCGGAGCCGAGCACGTGAAC AATAGCTACGAGTGCGACATCCCCATCGGCGCTGGAATCTGCGCCAGCTACCAGACACAG ACAAACAGCCCTCGGAGAGCCAGAAGCGTGGCCAGCCAGAGCATCATTGCCTACACAATG TCTCTGGGCGCCGAGAACAGCGTGGCCTACTCCAACAACTCTATCGCTATCCCCACCAAC TTCACCATCAGCGTGACCACAGAGATCCTGCCTGTGTCCATGACCAAGACCAGCGTGGAC TGCACCATGTACATCTGCGGCGATTCCACCGAGTGCTCCAACCTGCTGCTGCAGTACGGC AGCTTCTGCACCCAGCTGAATAGAGCCCTGACAGGGATCGCCGTGGAACAGGACAAGAAC ACCCAAGAGGTGTTCGCCCAAGTGAAGCAGATCTACAAGACCCCTCCTATCAAGGACTTC GGCGGCTTCAATTTCAGCCAGATTCTGCCCGATCCTAGCAAGCCCAGCAAGCGGAGCTTC ATCGAGGACCTGCTGTTCAACAAAGTGACACTGGCCGACGCCGGCTTCATCAAGCAGTAT GGCGATTGTCTGGGCGACATTGCCGCCAGGGATCTGATTTGCGCCCAGAAGTTTAACGGA CTGACAGTGCTGCCTCCTCTGCTGACCGATGAGATGATCGCCCAGTACACATCTGCCCTG CTGGCCGGCACAATCACAAGCGGCTGGACATTTGGAGCAGGCGCCGCTCTGCAGATCCCC TTTGCTATGCAGATGGCCTACCGGTTCAACGGCATCGGAGTGACCCAGAATGTGCTGTAC GAGAACCAGAAGCTGATCGCCAACCAGTTCAACAGCGCCATCGGCAAGATCCAGGACAGC CTGAGCAGCACAGCAAGCGCCCTGGGAAAGCTGCAGGACGTGGTCAACCAGAATGCCCAG GCACTGAACACCCTGGTCAAGCAGCTGTCCTCCAACTTCGGCGCCATCAGCTCTGTGCTG AACGATATCCTGAGCAGACTGGACCCTCCTGAGGCCGAGGTGCAGATCGACAGACTGATC ACAGGCAGACTGCAGAGCCTCCAGACATACGTGACCCAGCAGCTGATCAGAGCCGCCGAG ATTAGAGCCTCTGCCAATCTGGCCGCCACCAAGATGTCTGAGTGTGTGCTGGGCCAGAGC AAGAGAGTGGACTTTTGCGGCAAGGGCTACCACCTGATGAGCTTCCCTCAGTCTGCCCCT CACGGCGTGGTGTTTCTGCACGTGACATATGTGCCCGCTCAAGAGAAGAATTTCACCACC GCTCCAGCCATCTGCCACGACGGCAAAGCCCACTTTCCTAGAGAAGGCGTGTTCGTGTCC AACGGCACCCATTGGTTCGTGACACAGCGGAACTTCTACGAGCCCCAGATCATCACCACC GACAACACCTTCGTGTCTGGCAACTGCGACGTCGTGATCGGCATTGTGAACAATACCGTG TACGACCCTCTGCAGCCCGAGCTGGACAGCTTCAAAGAGGAACTGGACAAGTACTTTAAG AACCACACAAGCCCCGACGTGGACCTGGGCGATATCAGCGGAATCAATGCCAGCGTCGTG AACATCCAGAAAGAGATCGACCGGCTGAACGAGGTGGCCAAGAATCTGAACGAGAGCCTG ATCGACCTGCAAGAACTGGGGAAGTACGAGCAGTACATCAAGTGGCCCTGGTACATCTGG CTGGGCTTTATCGCCGGACTGATTGCCATCGTGATGGTCACAATCATGCTGTGTTGCATG ACCAGCTGCTGTAGCTGCCTGAAGGGCTGTTGTAGCTGTGGCAGCTGCTGCAAGTTCGAC GAGGACGATTCTGAGCCCGTGCTGAAGGGCGTGAAACTGCACTACACATGATGACTCGAG CTGGTACTGCATGCACGCAATGCTAGCTGCCCCTTTCCCGTCCTGGGTACCCCGAGTCTC CCCCGACCTCGGGTCCCAGGTATGCTCCCACCTCCACCTGCCCCACTCACCACCTCTGCT AGTTCCAGACACCTCCCAAGCACGCAGCAATGCAGCTCAAAACGCTTAGCCTAGCCACAC CCCCACGGGAAACAGCAGTGATTAACCTTTAGCAATAAACGAAAGTTTAACTAAGCTATA CTAACCCCAGGGTTGGTCAATTTCGTGCCAGCCACACCCTGGAGCTAGCAAAAAAAAAAA AAAAAAAAAAAAAAAAAAAAA
@P_J_Buckhaults - Phillip Buckhaults
democratize science.

@P_J_Buckhaults - Phillip Buckhaults
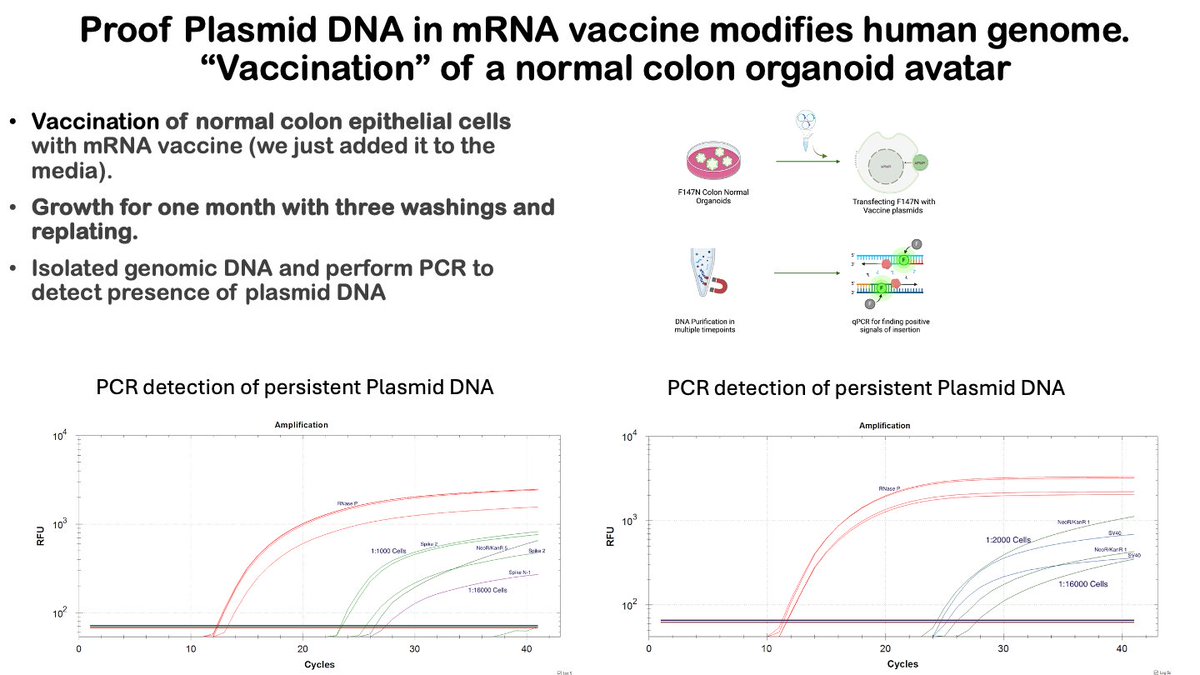
the plasmid DNA that is contained within mRNA vaccines can integrate into the genome of normal cells. i knew this could happen, but some were unconvinced, so we took the time to prove this in the lab. we grow normal human epithelial stem cells in my lab. its part of our normal job (cancer research). they are called organoids. these are not cancer cells, they are just the normal stem cells that make up the human colon. we "vaccinated" some of these normal cells and grew them for a month and saw pieces of the plasmid DNA persisting in the genomic DNA of the "vaccinated" cells. we detected the plasmid DNA with our qPCR protocol that was posted to X several months ago. this experiments was done mainly for the people who were paid to publicly ridicule this idea (and slander my reputation). most of these people are mutually blocked now, so pass around to anyone who needs to see it. maybe @DrPaulOffit or that rude gorsky dude would like to see it. IDK. this does not mean that the integration is happening in real vaccinated humans (those experiments are ongoing) but it does prove that the DNA can get into normal cells just fine, as i told everyone a year ago. its a small point and maybe looks like i am being petty, but the disparaging remarks made about me here and in the press really annoyed me so i thought it best to answer with a pipette. this experiment costs several thousands of dollars, whereas the reputation slander cost the saboteurs nothing. its asymmetric reputation warfare of sorts. anyway, here it is. peace.
@P_J_Buckhaults - Phillip Buckhaults
In my view, this kind of assay (pour stuff onto normal human organoids) would be a really good way to check all kinds of things we consume for their mutagenic potential. we could see into the future by decades by identifying things that mutate DNA. think of this as an AMES test in human cells. this would be a smart thing for the US FDA to start doing in house.

@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
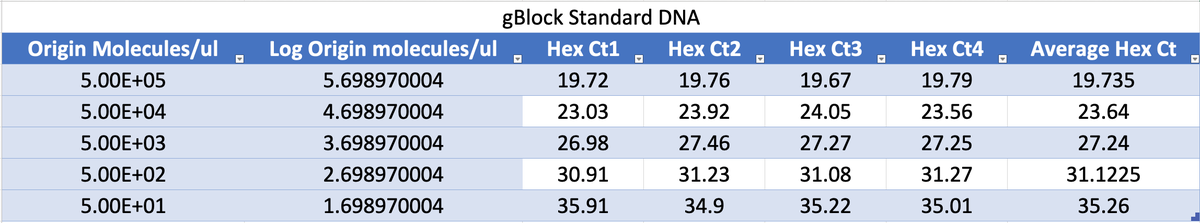
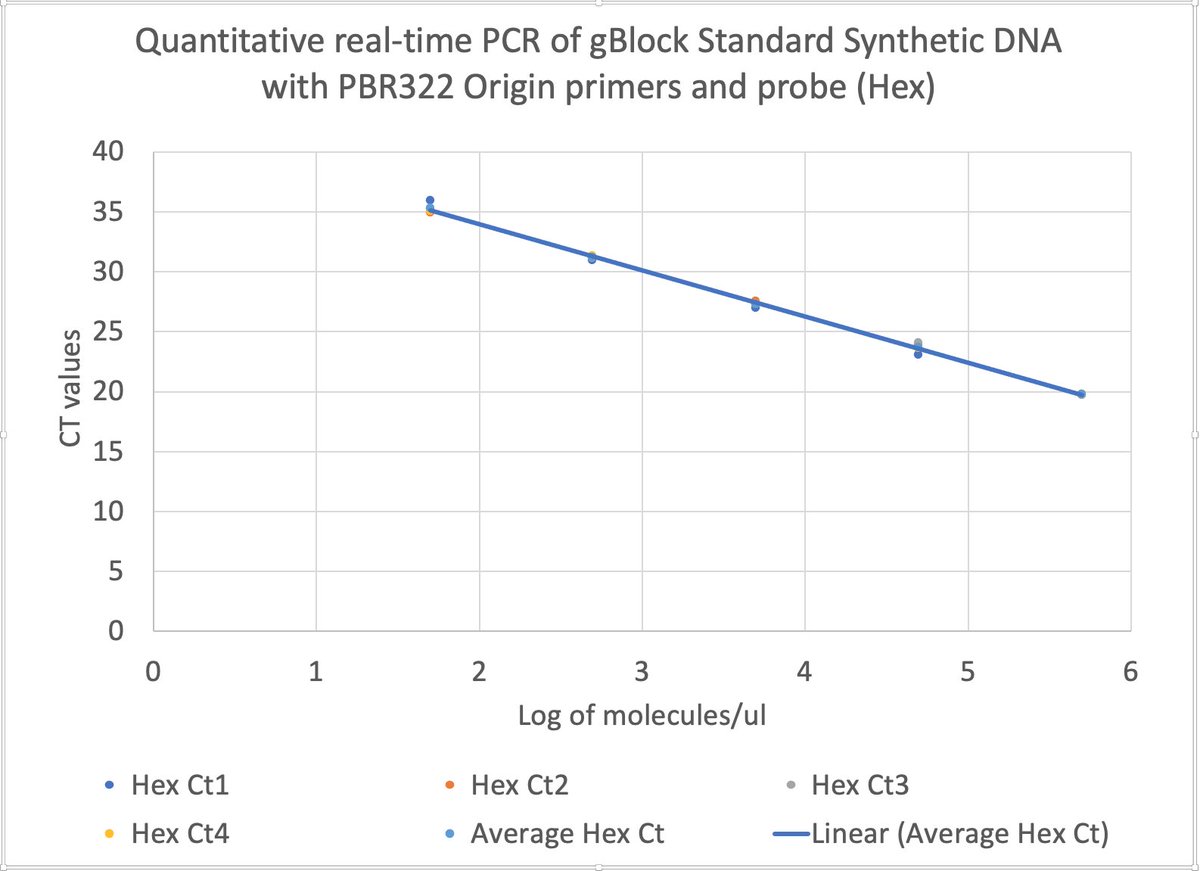
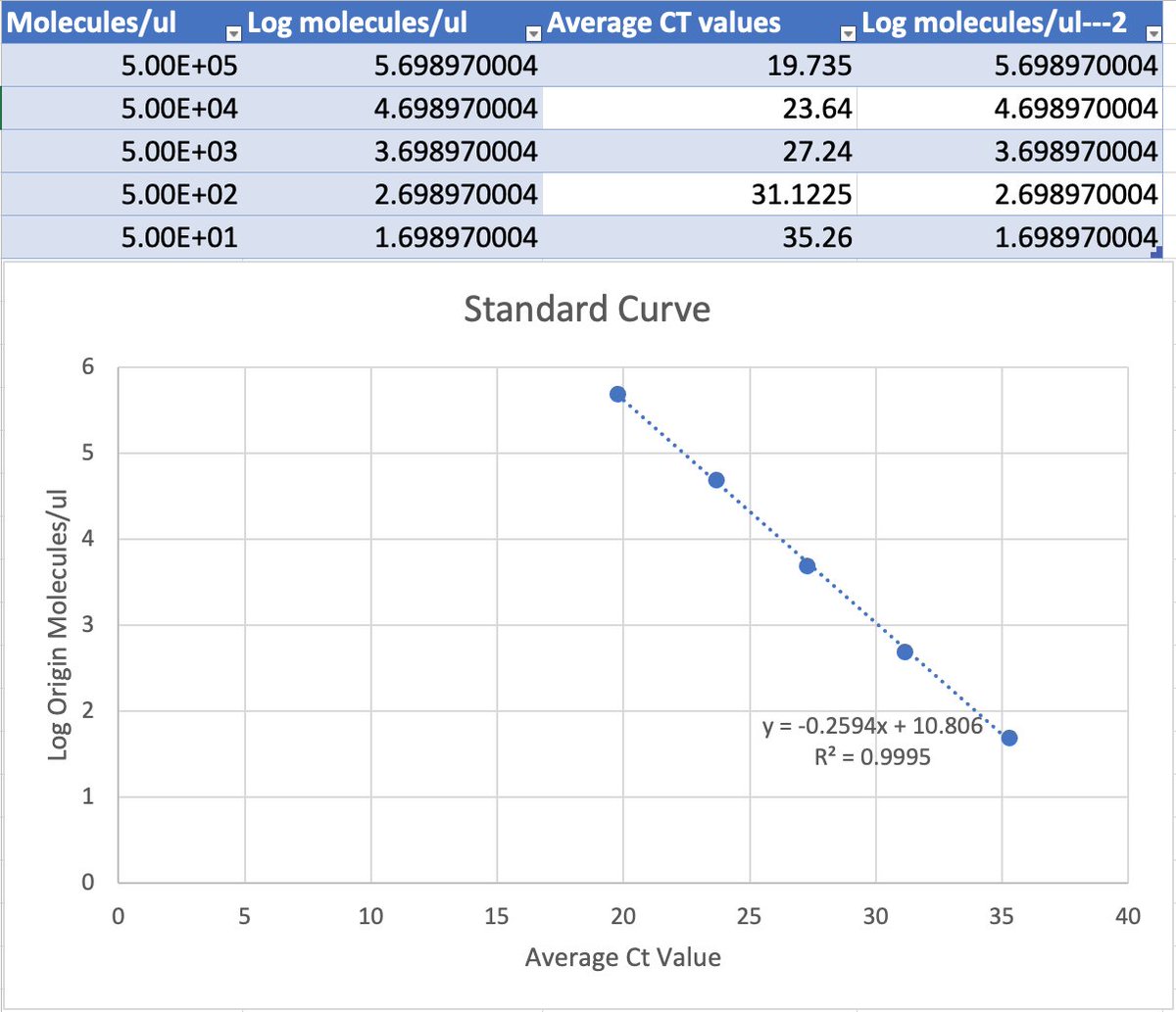
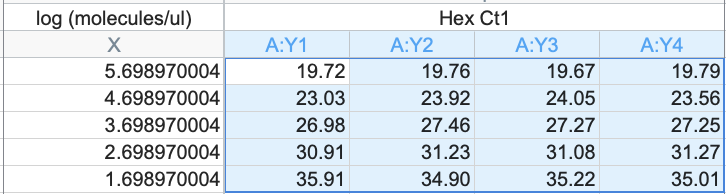
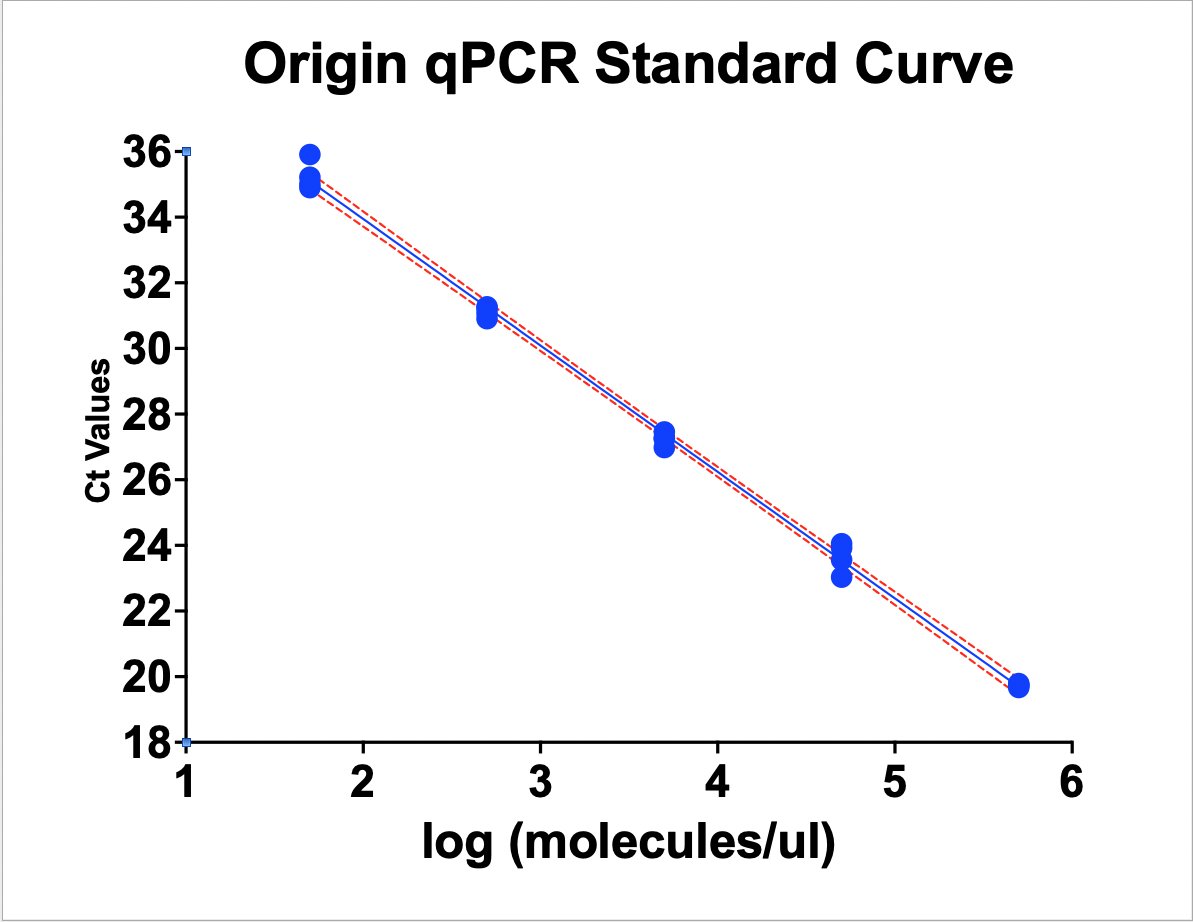
this works out to be about 2.5 million molecules per microliter in raw vaccine, or 675-774 million molecules per dose (300ul). thats a lot of chances for DNA to transfect and integrate. multiply this by the number of arms that it has been put into world wide and i guarantee you there has been genome modification. only question is how frequent and did it every do anything bad.
@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
@Kevin_McKernan @US_FDA you should revisit this topic. have someone look for the presence or absence of plasmid DNA integrated into the genomes of long live stem cells of vaccinated individuals. this is not a crazy idea.

@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
Plasmid DNA is a low but non-zero risk for insertional mutagenesis of normal stem cells. IMO, the mRNA vaccination platform will be at its best when it is completely devoid of plasmid DNA pieces (yes, i know others whom i respect will disagree with me on this point). Regarding the DNA subject.... I have transfected billions of cells with DNA fragments during my career and i have an informed intuitive sense (based on data and experience) for the in vivo cancer risks posed by low level DNA contamination in these products. I suspect the risk is similar to smoking and lung cancer or excessive sun exposure and melanoma. the risk is concerning enough to study and minimize but not so high to frighten people who have already been vaccinated; that's just not ethical or helpful. Now, we can quantify this risk by one of two methods. The first method is wait for 30 years and count the number of cases that show up with DNA plasmid as a driver event. The second method is to perform single cell sequencing of normal cells (that are PCR positive) and count the frequencies of integrations into cancer driver loci. The latter method costs a bit of money but is able to look into the future and assess risk before it comes to clinical presentation. Either way, we will eventually know what the risks are, and it may end up being so low that we would not care (the benefits eclipse the risks) or it may be high enough that we should change the regulatory limits for allowable DNA. I vote for using the second method but that's just me because the technology is what our lab does (to a man with a hammer everything looks like a nail). Others may disagree and feel we should just let the epidemiology tell us over the next decade. Mine is not an ant-vax stance or an anti-pharma stance or an anti-mRNA vax stance, but rather is a pro-person stance. We did not check these things initially probably because nobody thought of it (and the house was on fire and most people were sincerely scared). It is my hope to cause policy makers and industry experts to think about it now and engage in careful genome-scarring safety studies using single cell sequencing technology, to accurately quantify these risks going forward. I do not see any "fault" here. I simply see a better way to do things going forward.

@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
Dear @US_FDA and @pfizer Phillip Buckhaults, PhD. here. Cancer genejock faulty at the university of South Carolina. I have recently analyzed a couple of lots of vaccine by nanopore sequencing of all the DNA in the vax, and by qPCR (DNA pcr, not reverse-transcriptase pcr) using primers directed at vector backbone sequences. (credit below) long story short… the Pfizer mRNA vaccine is contaminated with the plasmid DNA vector that was used as the template for in vitro transcription reaction. There are about 2 billion copies of the fragment containing the origin of replication, and from nanopore sequence analysis, there is probably 50-100 times that many pieces of plasmid DNA derived from the entire vector. This means each shot has about 200 billion pieces of genomic DNA encapsulated in the lipid nanoparticle. This DNA could be the cause of some of the rare side effects. it could integrate into the genomes of transfected cells. if any lipid nanoparticle complexes find their way to stem cell niches there is a hazard for genome modification of long lived somatic cells, which could cause autoimmune attack toward that tissue or (rarely) cancer, depending on the piece of DNA and site of integration. I think there is a serious hazard of immune based illnesses that should be investigated at the regulatory level. The amounts of DNA in each vax is close to allowable limits. if it is over, its not too much over. The trouble is, i think these limits may have been established for naked DNA present in a recombinant protein vax (or an attenuated virus produced in cell culture) not for DNA encapsulated in a transfection liposome. Even if the vax is within regulatory limits, the regulatory limits may need to be re-visited in light of the propensity of lipid-nanoparticle-DNA to transform human stem cells. IMO it would be wise to check all lots/batches to see what the range of DNA contamination is, and it would be wise to check tissue samples from select people for evidence of presence of this plasmid integrating into stem cell genomes. this experiment could yield valuable negative information as well, by proving that integration never happens. either way, the public will be served by such analyses of somatic genome integrity plus and minus vaccination. credit to @Kevin_McKernan who first had the good sense and technical skills to sequence vaccine and then to design elegant qPCR assays to detect this stuff.
@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
Dear @US_FDA and @pfizer Phillip Buckhaults, PhD. here. Cancer genejock faulty at the university of South Carolina. I have recently analyzed a couple of lots of vaccine by nanopore sequencing of all the DNA in the vax, and by qPCR (DNA pcr, not reverse-transcriptase pcr) using primers directed at vector backbone sequences. (credit below) long story short… the Pfizer mRNA vaccine is contaminated with the plasmid DNA vector that was used as the template for in vitro transcription reaction. There are about 2 billion copies of the fragment containing the origin of replication, and from nanopore sequence analysis, there is probably 50-100 times that many pieces of plasmid DNA derived from the entire vector. This means each shot has about 200 billion pieces of genomic DNA encapsulated in the lipid nanoparticle. This DNA could be the cause of some of the rare side effects. it could integrate into the genomes of transfected cells. if any lipid nanoparticle complexes find their way to stem cell niches there is a hazard for genome modification of long lived somatic cells, which could cause autoimmune attack toward that tissue or (rarely) cancer, depending on the piece of DNA and site of integration. I think there is a serious hazard of immune based illnesses that should be investigated at the regulatory level. The amounts of DNA in each vax is close to allowable limits. if it is over, its not too much over. The trouble is, i think these limits may have been established for naked DNA present in a recombinant protein vax (or an attenuated virus produced in cell culture) not for DNA encapsulated in a transfection liposome. Even if the vax is within regulatory limits, the regulatory limits may need to be re-visited in light of the propensity of lipid-nanoparticle-DNA to transform human stem cells. IMO it would be wise to check all lots/batches to see what the range of DNA contamination is, and it would be wise to check tissue samples from select people for evidence of presence of this plasmid integrating into stem cell genomes. this experiment could yield valuable negative information as well, by proving that integration never happens. either way, the public will be served by such analyses of somatic genome integrity plus and minus vaccination. credit to @Kevin_McKernan who first had the good sense and technical skills to sequence vaccine and then to design elegant qPCR assays to detect this stuff.
@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
@US_FDA @pfizer Faulty <> Faculty. lol.

@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
Latest nanopore sequencing of all DNA found in @pfizer mRNA vaccine. there is quite a large number of little pieces of doublestranded DNA fragments derived from the vector used to make the mRNA. @US_FDA guidelines were set using studies of naked DNA injected into animals and…

@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
Latest nanopore sequencing of all DNA found in @pfizer mRNA vaccine. there is quite a large number of little pieces of doublestranded DNA fragments derived from the vector used to make the mRNA. @US_FDA guidelines were set using studies of naked DNA injected into animals and seeing no ill effects. this vaccine is not naked DNA, but DNA encapsulated in a lipid transfection particle. it is protected until it is efficiently delivered into cells. the mutagenic potential of this DNA is lower than retroviruses but higher than naked DNA. someone should perform a careful single celll sequencing study of vaccinated people and document the absence (or presence) of integrated pieces of DNA. the vaccine has saved a lot of people from the graveyard, and it can be made even better by getting this DNA out of the manufacturing process. @US_FDA please insist that @pfizer conduct long term studies of somatic cells genome integrity after vaccination. i think this is important.
@EdV1694 - Robert Kogon (former "Esprit de Voltaire")
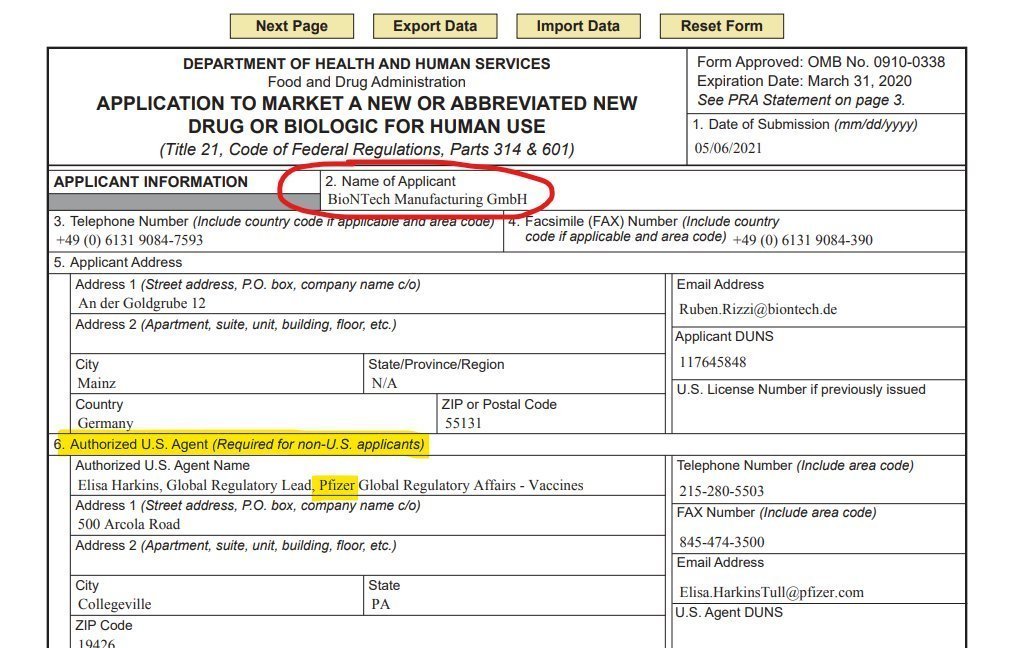
@P_J_Buckhaults @pfizer @US_FDA 1/This is obviously important issue. But Pfizer is not the license-holder, the German company BioNTech is, And, btw, the problem of DNA contamination has long been known in Germany. Sponsor of & responsible for such studies is always BIONTECH. E.g. https://classic.clinicaltrials.gov/ct2/show/NCT04368728

@EdV1694 - Robert Kogon (former "Esprit de Voltaire")
@P_J_Buckhaults @pfizer @US_FDA 3/ Pfizer merely acted as BioNTech's (legally required) US agent and collaborator in the authorization process. See below.
@EdV1694 - Robert Kogon (former "Esprit de Voltaire")
@P_J_Buckhaults @pfizer @US_FDA 4/ No offense. But why is the entire opposition persistently barking up the wrong tree and typically not even mentioning BioNTech? It's very harmful and means that BioNTech, the legal manufacturer, license holder and legally responsible party, will never be held accountable.
@EdV1694 - Robert Kogon (former "Esprit de Voltaire")
@EdV1694 - Robert Kogon (former "Esprit de Voltaire")
@P_J_Buckhaults @pfizer @US_FDA P.S. Or here at Brownstone... https://brownstone.org/articles/read-the-label-its-biontechs-vaccine-not-pfizers/

@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
Omicrons (B.1.1.529) most recent common ancestor is AV.1. Relative to this MRCA, Omicron has 25 nonsynonymous and 1 synonymous mutation in Spike, and 13 nonsynonymous and 6 synonymous mutations in the entire rest of the genome. This is very strange & I don't understand.
@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
I have re-done the alignments and counting up of mutations, and it's even weirder than i first said.
@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
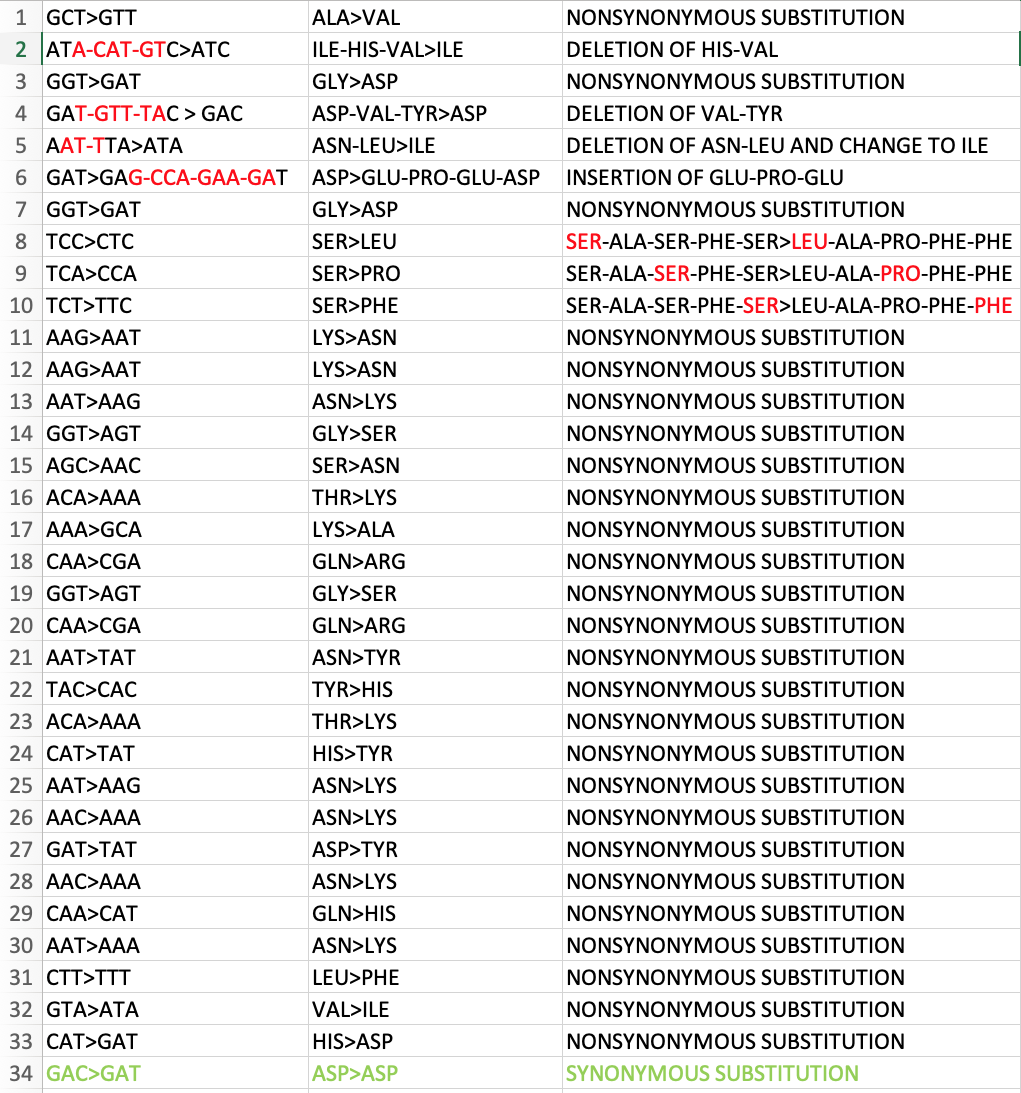
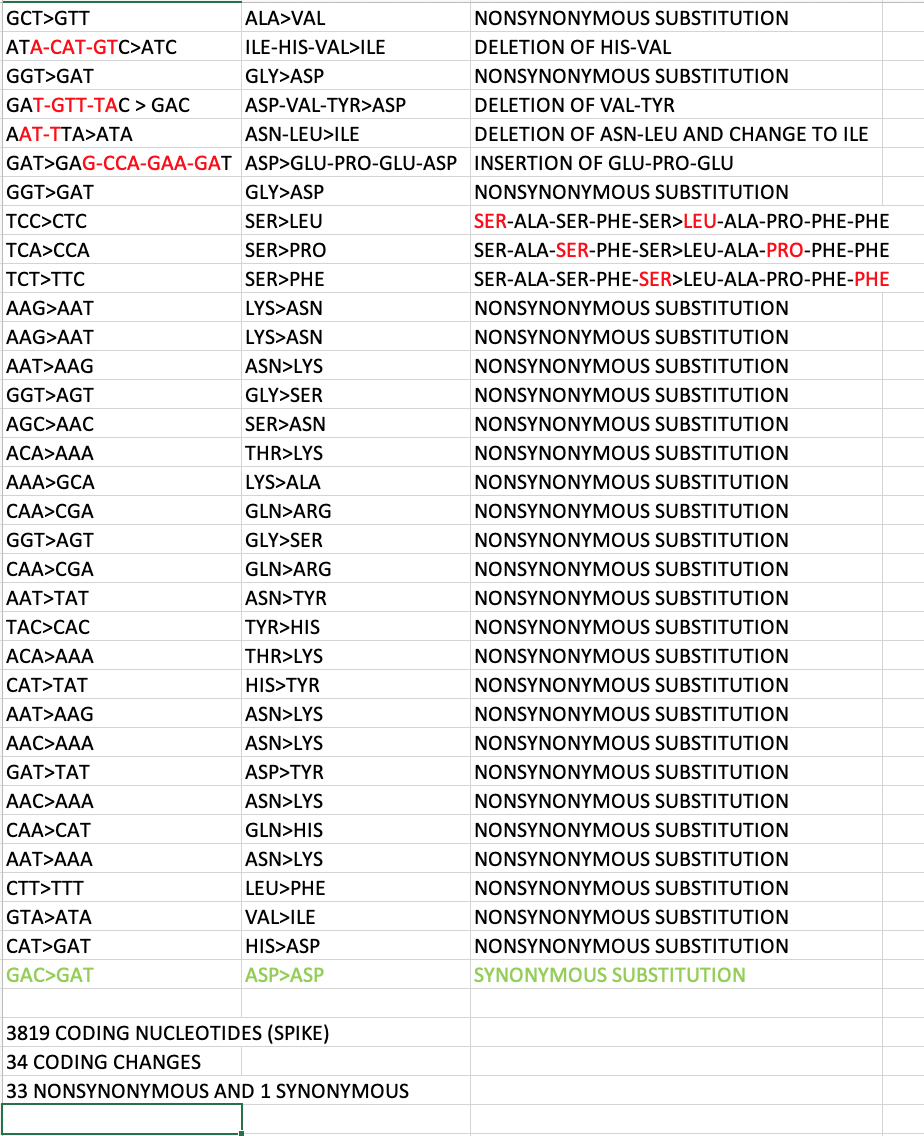
34 discontinuous changes (that are likely independent events). only one of them is silent. the rest change amino acid. see attached pictures. to quote @K_G_Andersen this is inconsistent with expectations from standard evolutionary theory. https://t.co/y1DktIAogr
@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
this is just bonkers. there should be a butt-tonne of other silent passenger mutations hitching a ride with nonsynonymous changes under positive selective pressure. i am not invoking a sinister lab engineering event here, just something important that i do not know.
@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
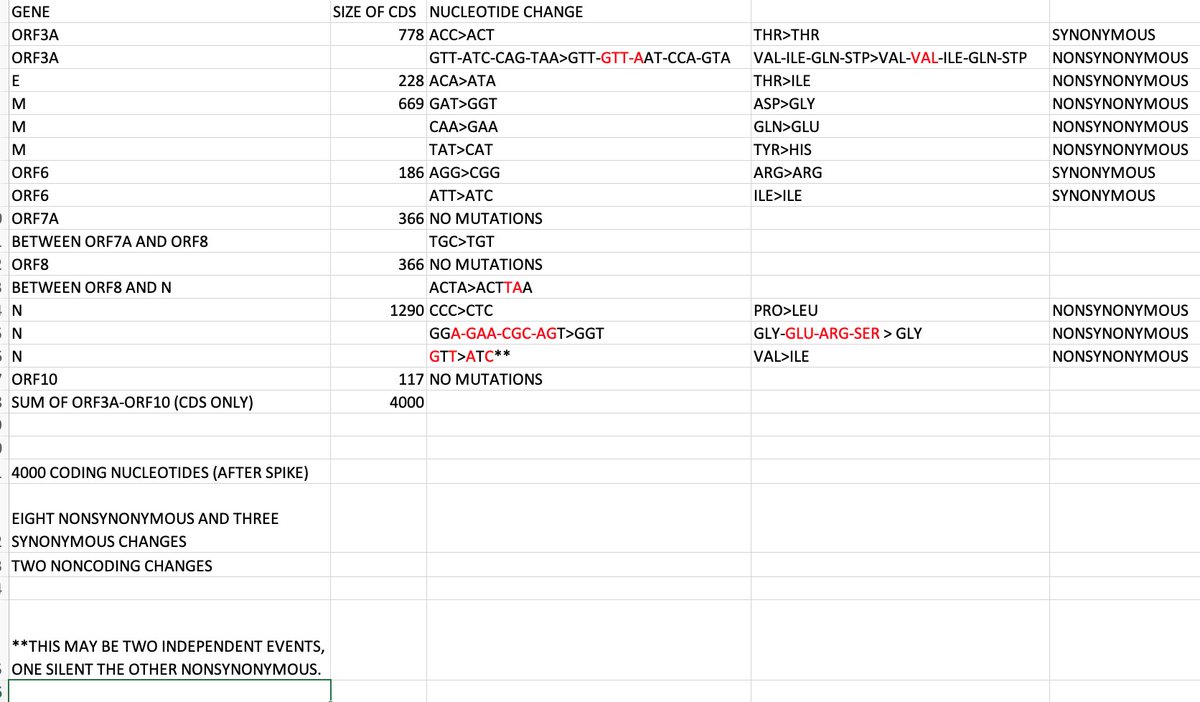
Spike is 3819 coding nucleotides. It has 33 NS and 1 S changes. The neighboring ORF3A-E-M-ORF6-ORF7A-ORF8-N-ORF10 genes have 4000 coding nucleotides. They have eight NS and three S changes. Spike is smaller and has four times the NS changes. https://t.co/emqNEgp0FC
@P_J_Buckhaults - Phillip J. Buckhaults, Ph.D.
positive selection on spike should drag a bunch of synonymous changes in the neighborhood along for the ride. unless spike can evolve separately from the rest of the genome.