reSee.it - Tweets Saved By @stevenemassey

@stevenemassey - Steve Massey
Zhengli Shi of the WIV constructed a dangerous Gain of Function MERS chimera that contaminated pre-pandemic rice sequence datasets from Wuhan Potential violation of the Biological Weapons Convention NIAID + USAID provided funding Parallels with SARS2 🧵 #DRASTIC https://t.co/mrFgW2eSR3

@stevenemassey - Steve Massey
The Zhang group of Fudan University have identified and validated two A-B intermediate SARS2 genomes from the early pandemic This provides a key to understanding the origin of COVID19 🧵
@stevenemassey - Steve Massey
2/ In their new paper, the Zhang group sequence 343 new SARS2 genomes from the early pandemic (sampled up to Oct 2020). The genomes were obtained from COVID19 patients in the Shanghai Public Health Center https://academic.oup.com/ve/advance-article/doi/10.1093/ve/veae020/7619252?login=false
@stevenemassey - Steve Massey
3/ Importantly, they identify two SARS2 genomes intermediate between lineage A and lineage B These were validated using two methods, RT-PCR (Sanger sequencing), and Next Generation Sequencing (NGS). @jbloom_lab verified the sequencing depth on one (high)
@stevenemassey - Steve Massey
3/ What is an A-B intermediate genome and why is it important ? Lineages A and B were the first major lineages to emerge during the early pandemic. They are only separated by two mutations, at positions 8782 and 28144
@stevenemassey - Steve Massey
4/ Lineage A is T8782 /C28144 (T/C) while lineage B is C8782/T28144 (C/T) The closest related bat CoVs are T/C implying A is ancestral A and B interconverted via a single mutation, either via C8782 / C28144 (C/C) or T8782/T28144 (T/T)
@stevenemassey - Steve Massey
5/ The existence of either a T/T or C/C intermediate in the human population would indicate that this interconversion occurred after SARS2 entered the human population, supporting a single introduction This is why intermediates are key to understanding the origin of the pandemic
@stevenemassey - Steve Massey
6/ The two T/T intermediate genomes sequenced by the Zhang group from patients infected in Henan and Shanghai and hospitalized on Feb 4th and Feb 8th 2020 respectively
@stevenemassey - Steve Massey
7/ These are related to 7 T/T genomes in the db: 2 from Wuhan, 4 from Singapore and 1 from the UAE Notably, 3 of these are identical to the two new T/T intermediates sequenced by the Zhang group, and 3 more only differed by a single SNV
@stevenemassey - Steve Massey
8/ The widely cited Pekar et al (2022) posited that there were two separate introductions of lineage A and B, in the Huanan Seafood Market (HSM) A major plank of their thesis was the claimed absence of true intermediate sequences https://www.science.org/doi/10.1126/science.abp8337
@stevenemassey - Steve Massey
9/ However, @humblesci @Daoyu15 @ydeigin @quay_dr and myself previously showed that their exclusion criteria were flawed, and that several potential intermediates were improperly excluded by Pekar et al https://www.mdpi.com/2036-7481/14/1/33

@stevenemassey - Steve Massey
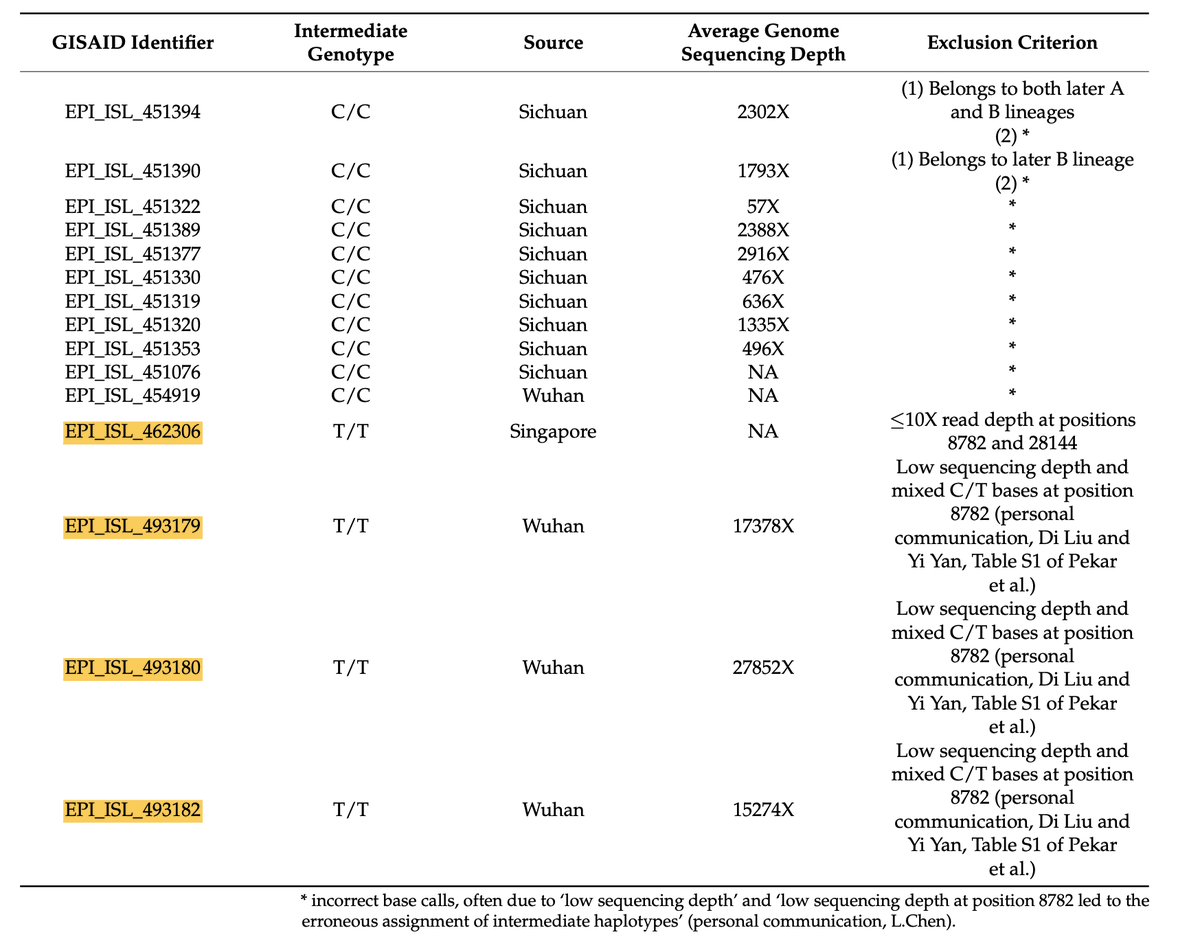
10/ This included 4 potential T/T intermediates, 3 of which were noted by the Zhang group (EPI_ISL_462306, EPI_ISL_493180 and EPI_ISL_493182) In our paper we argue all four were improperly excluded, on the basis of personal communications, and abitrary use of depth cutoffs
@stevenemassey - Steve Massey
11/ The fourth, EPI_ISL_493179, was not mentioned by the Zhang group, but was from Wuhan and part of the same study that generated EPI_ISL_493180 and EPI_ISL_493182) It differs from Hu-1 at C8782T, T13402G
@stevenemassey - Steve Massey
12/ In addition, with @WashburneAlex we identified an additional T/T intermediate was not considered at all by Pekar et al (or the Zhang group) This was OM065349 (Genbank Accession), sampled in Lu'an, Anhui on 30 Jan 2020 from a 53 yr old female https://www.biorxiv.org/content/10.1101/2022.10.10.511625v1

@stevenemassey - Steve Massey
13/ This genome is identical to the 2 new T/T intermediates from the Zhang group In total, there are 4 intermediates in the db that differ from Hu-1 only at C8782T (that gives the T/T genotype) and are identical to the 2 new T/T intermediates from Zhang et al
@stevenemassey - Steve Massey
14/ So, there are 6 identical T/T intermediates, sampled from a variety of locations in and outside China, early in pandemic
@stevenemassey - Steve Massey
15/ Why is this important ? The existence of T/T intermediates in the human population indicates a single introduction of SARS2 1) This confirms that lineage A is ancestral This is because A is T/C, the same as the closest related bat CoVs
@stevenemassey - Steve Massey
16/ 2) This excludes the HSM as source of the spillover This is because the genomes sequenced from the HSM were almost all B, which is derived, as opposed to A, which is ancestral
@stevenemassey - Steve Massey
17/ 3) This indicates a date of emergence of no later than ~ Oct 2019, as per Kumar et al This is based on the number of mutations needed (3) to get to proCoV2 from the lineage B reference sequence (Hu-1) https://academic.oup.com/mbe/article/38/8/3046/6257226
@stevenemassey - Steve Massey
18/ Finally, a further potential clue to the origin of the pandemic is presented by our preprint by @humblesci @Daoyu15 @BiophysicsFL @ydeigin @quay_dr and myself characterizing a MERS-related infectious clone from Wuhan 2019 that has undergone apparent GOF experimentation
@stevenemassey - Steve Massey
19/ It was recently turned down from a journal for non-scientific reasons, in an apparent failure of nerve on the part of reviewers and editor https://www.biorxiv.org/content/10.1101/2023.02.12.528210v2

@stevenemassey - Steve Massey
20/ Are there any journal editors out there brave enough to give our manuscript a fair hearing ? https://t.co/AS9YRH7hRb
@stevenemassey - Steve Massey
21/ @threadreaderapp unroll