reSee.it - Tweets Saved By @tommy_cleary

@tommy_cleary - Tommy Cleary
Holmes attempted <
@tommy_cleary - Tommy Cleary
One more before I put the roast on for Australia Day dinner...
@GrahamPerrettMP is my local Federal MP and he has helped in the past, but last time I wrote to him he replied that I should check the Queensland State Library for more details...perhaps I should check back with him again too.
These issues of how to handle the dangerous side of science have been a problem since at least Iraq's @UN biological weapons inspections...with discussions of Mustard brought to the table by @R_H_Ebright thank you, @INTERPOL_CBRNE questions are important.
H/t @CharlesRixey @Ayjchan @Globalbiosec
Holmes attempted <
@tommy_cleary - Tommy Cleary
Seeking No14 ORF8 omission...with some healthy distraction from @breakfast_dogs @harishseshadri2 @gdemaneuf
about the truisms of love...and knowing at all. @Rebecca21951651 @emilyakopp @a_kruschke @Ayjchan @VBruttel @BillyBostickson
Back to the data set.
Holmes attempted <

@tommy_cleary - Tommy Cleary
The question of //Pathos// has disturbed the search for the next missing part of this data set.
Never a better reason to interrupt seeking is finding a question linked to the heart.
Knowing love is a perennial concern.
To leave souls behind has a sharp gravitas.
Back to the data.
In 2023 Holmes attempted <
@tommy_cleary - Tommy Cleary
In 2023 Holmes attempted <
@tommy_cleary - Tommy Cleary
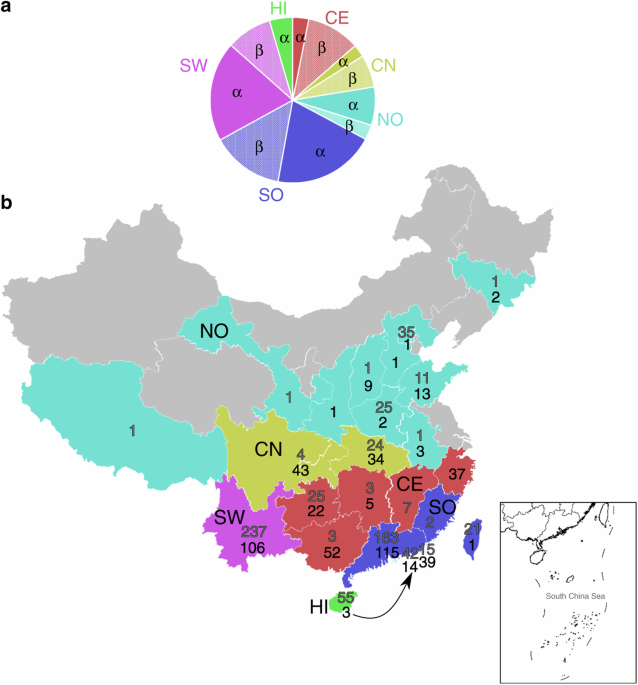
As science is very important...
https://journals.asm.org/doi/10.1128/jvi.01240-24methods
H/t @sciencecohen @hholdenthorp @ScienceMagazine
Holmes attempted
<

@tommy_cleary - Tommy Cleary
My application for SAGO at @WHO was rejected...but it was in volunteer capacity and so I simply continued to help where I can.
https://2012-2017.usaid.gov/sites/default/files/documents/2496/Combatting_Corruption_Among_Civil_Servants_-_Interdisciplinary_Perspectives_on_What_Works.pdf
My skill sets are listening...catching...and surprise...not simplicity
H/t @CharlesRixey
Umberto Eco said it well.
If it is too complicated, read more books.
But he wrote this type of thing in Italian, so don't see these ideas as complexity, see them as language.
Teaching a language takes time and repetition...about two years of immersion...or you can nowadays Gronk your way through?
The lived experience here is of an INCOMPLETE data set...so obviously I cannot fully explain the data...but you can join me on the journey.
Surprise!
Truth is important...but it takes a lot of listening to hear certain truths...trauma adds more layers of humanity and so our souls are stretched thinly as we listen to the person within the cyborg of text based embodiment twisting under the weight of the unknown...but knowable:
<

@tommy_cleary - Tommy Cleary
<

@tommy_cleary - Tommy Cleary
This is where I lost count!
Doh!
Next GI will have to start from here and be inserted into current tally.
GI 1769824546 restart count again here...and insert missing into tally
KISS Methods: basic GI series analysis this Xpost
GI is 1769824546 <
@tommy_cleary - Tommy Cleary
Censorship of the nature deployed in the case of COVID had some obvious negative effects...but some were not so bad. @BiosafetyNow
https://biosafetynow.substack.com/p/censoring-virology
It was nice and quiet.
The people censored had to find ways to reach out to each other...the phenomenology was that we had to look at what we were looking through.
It also builds a compassion for a data set you are auditing during verification and for your own findings...a health doubt...need to double check and have peers that are brutal not lazy.
Fixing this mess is going to be fun!
Some wisdom always comes from a moment of stupidity and reflection.
KISS Methods: basic GI series analysis this Xpost
GI 1769824482 <



@a_kruschke - A.Kruschke
@tommy_cleary @JAHawk94684 @MartinaSisters @mlperk1 @MonaRahalkar @R_H_Ebright @quay_dr @BiosafetyNow @syd_health @SystemsVirology @NLM_NIH @POTUS @Sydney_Uni @DrJBhattacharya @FloDebarre @institutpasteur @thackerpd @Rebecca21951651 @capitolsheila @BillyBostickson @breakfast_dogs @Globalbiosec @gdemaneuf @RdeMaistre @dasher8090 @COVIDSelect @GrahamPerrettMP @harishseshadri2 @emilyakopp @Ayjchan @VBruttel @CharlesRixey @sciencecohen @hholdenthorp @ScienceMagazine @WHO @reSeeIt save Thread