reSee.it - Tweets Saved By @ydeigin

@ydeigin - Yuri Deigin
🧵 WHAT REALLY HAPPENED IN DRASTIC Ok, we need to set the record straight about how the DRASTIC schism happened. And no, it’s not because of the Radical R, although many of us did want to change it once we realized Billy originally used it for his RAGE University years ago: https://t.co/RpuMYgKIGO
@ydeigin - Yuri Deigin
Needless to say, most DRASTIC members want nothing to do with such radical anarchist messaging which we never signed up to endorse: https://t.co/1PA74Tfwt0
@ydeigin - Yuri Deigin
But what precipitated our decision to separate from Billy was the revelation that he appropriated the group email access and his subsequent refusal to comply with group’s majority vote to share it with the group’s appointee.
@ydeigin - Yuri Deigin
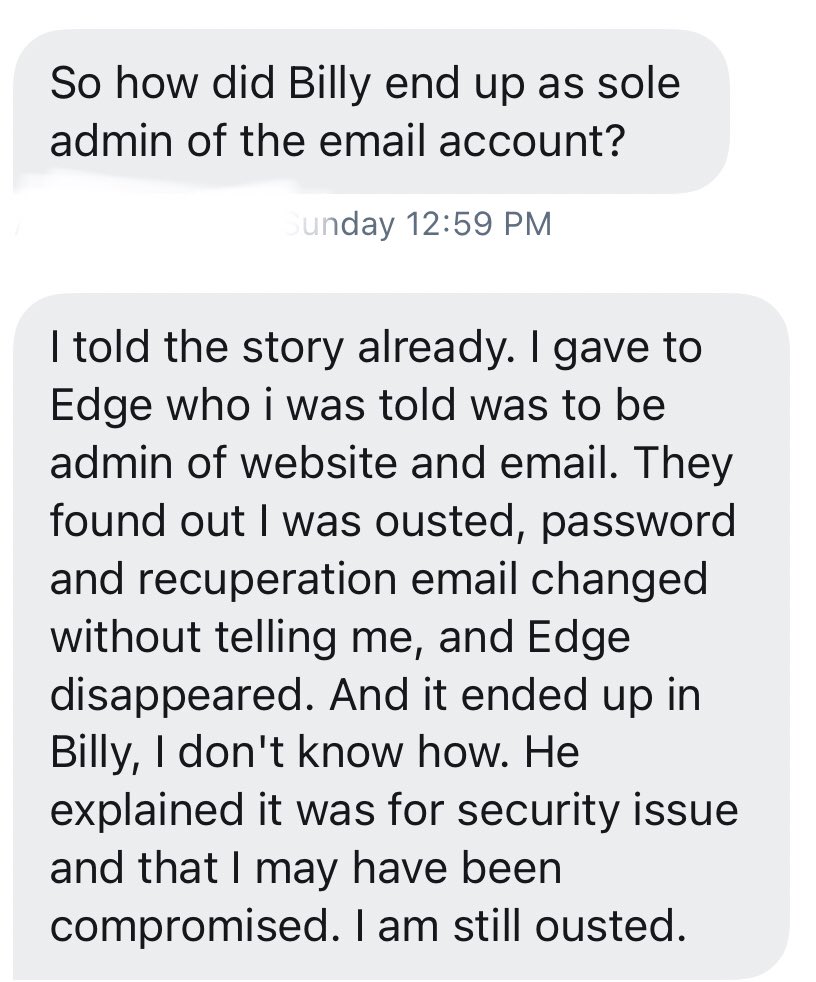
The original group email was created by @RdeMaistre, and he then shared its password with EdgeOfTheLeaf who was the website admin at the time. Later Rodolphe found himself locked out, and then learned it was Billy who changed not only the password but also the recovery email. https://t.co/pMr5dSqAd0
@ydeigin - Yuri Deigin
Billy then told Rodolphe a ridiculous story how EdgeOfTheLeaf deleted his Twitter account because of either getting cancer or fearing that he has compromised his anonymity. Surprisingly, Edge didn’t bother telling anyone else this or any other story before disappearing.
@ydeigin - Yuri Deigin
We found out about all this when @Rossana38510044 challenged Billy for his dictatorial behavior over access to journalists & filmmakers, but instead of agreeing to restore group access to email Billy insulted Rossana and then locked her and 3 more members out of the group.
@ydeigin - Yuri Deigin
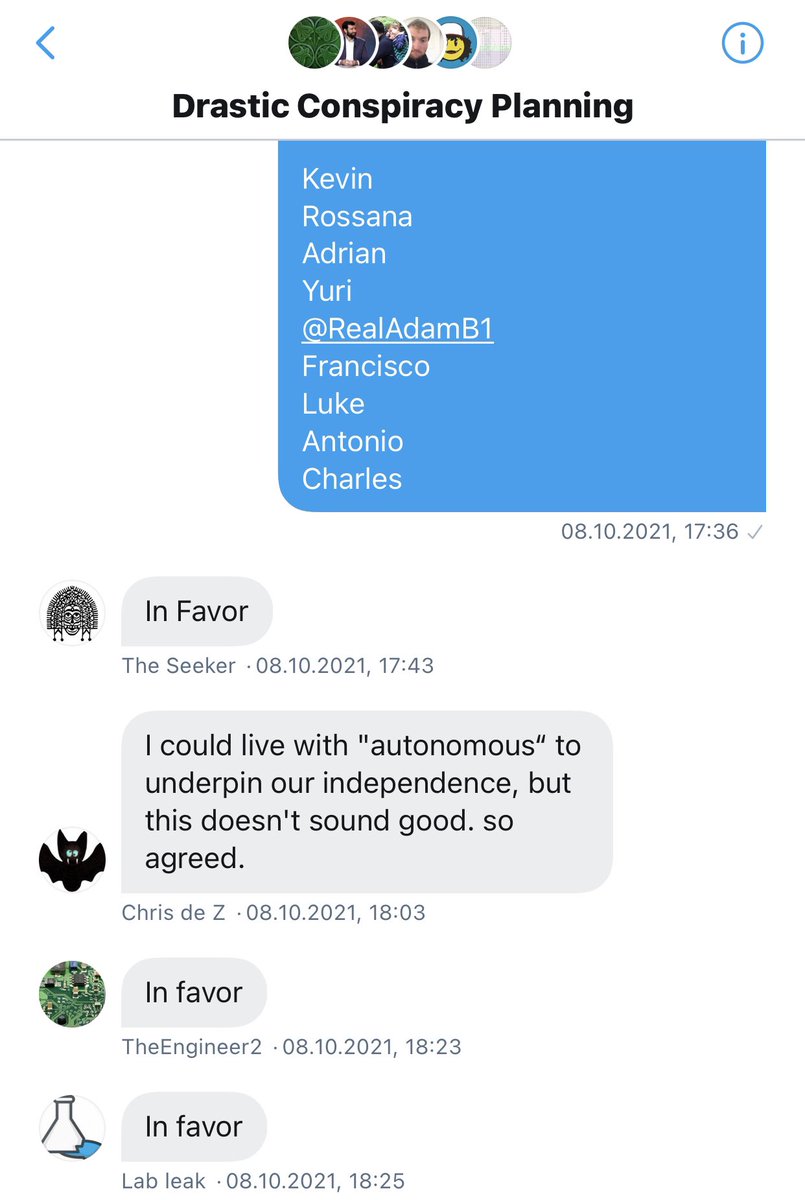
That was on October 2, and a multi-day crisis ensued where several members tried to convince Billy to share group email access but to no avail. The group then held a formal vote to compel Billy to share email access with at least me but he still refused. https://t.co/Y3XNmvh06S
@ydeigin - Yuri Deigin
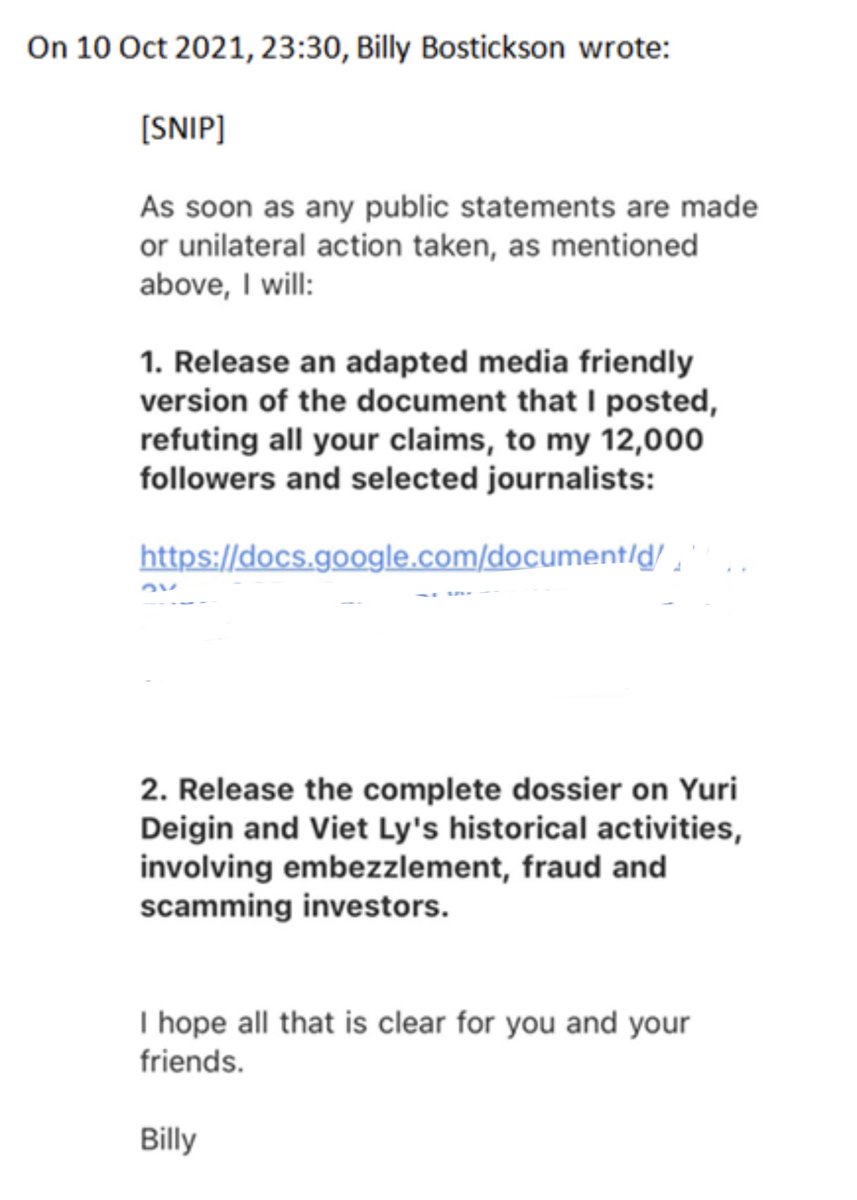
At that point we had no other choice but to create a new email and website, as Billy controlled both at the time. However, revealing his true terrorist nature, he threatened to engage in a slanderous smear campaign against me if we chose to make the new website and email public: https://t.co/hvEY3AfxiK
@ydeigin - Yuri Deigin
The group agreed that we do not negotiate with terrorists, and after we set up the new email and constructed the website, we made the announcement of distancing from Billy and his website.
@ydeigin - Yuri Deigin
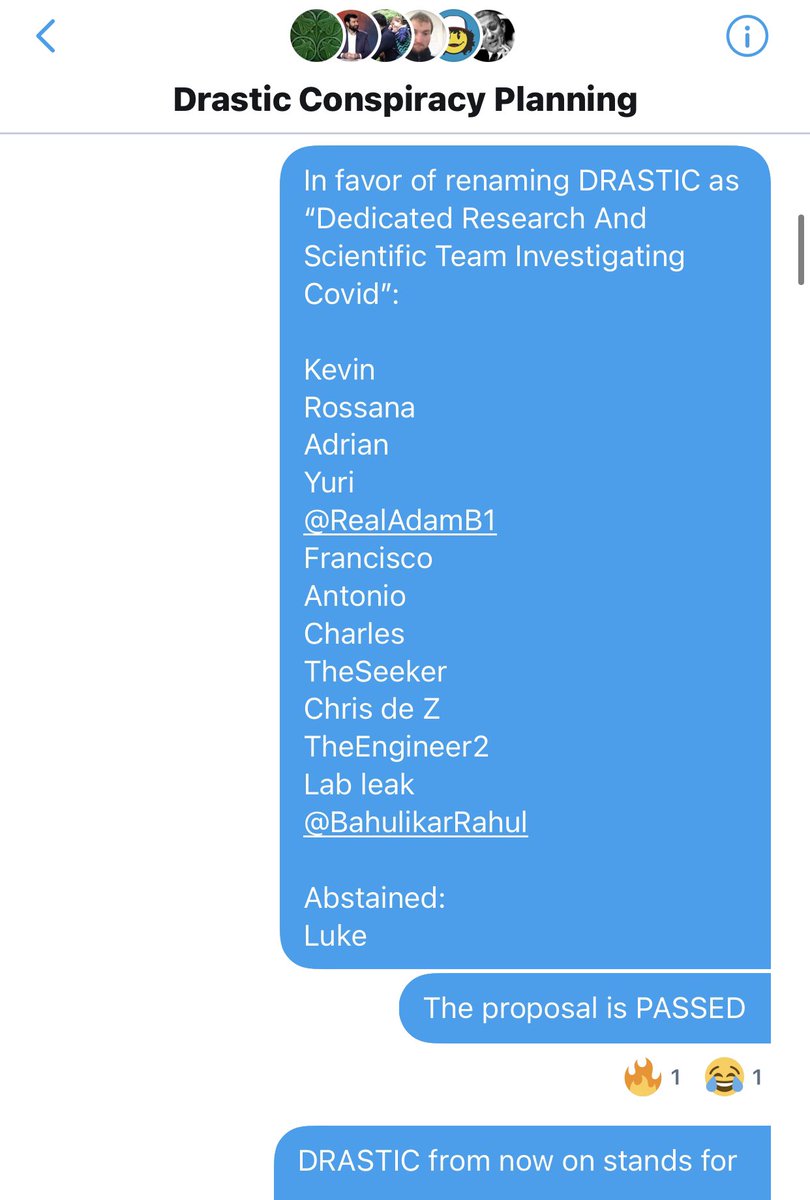
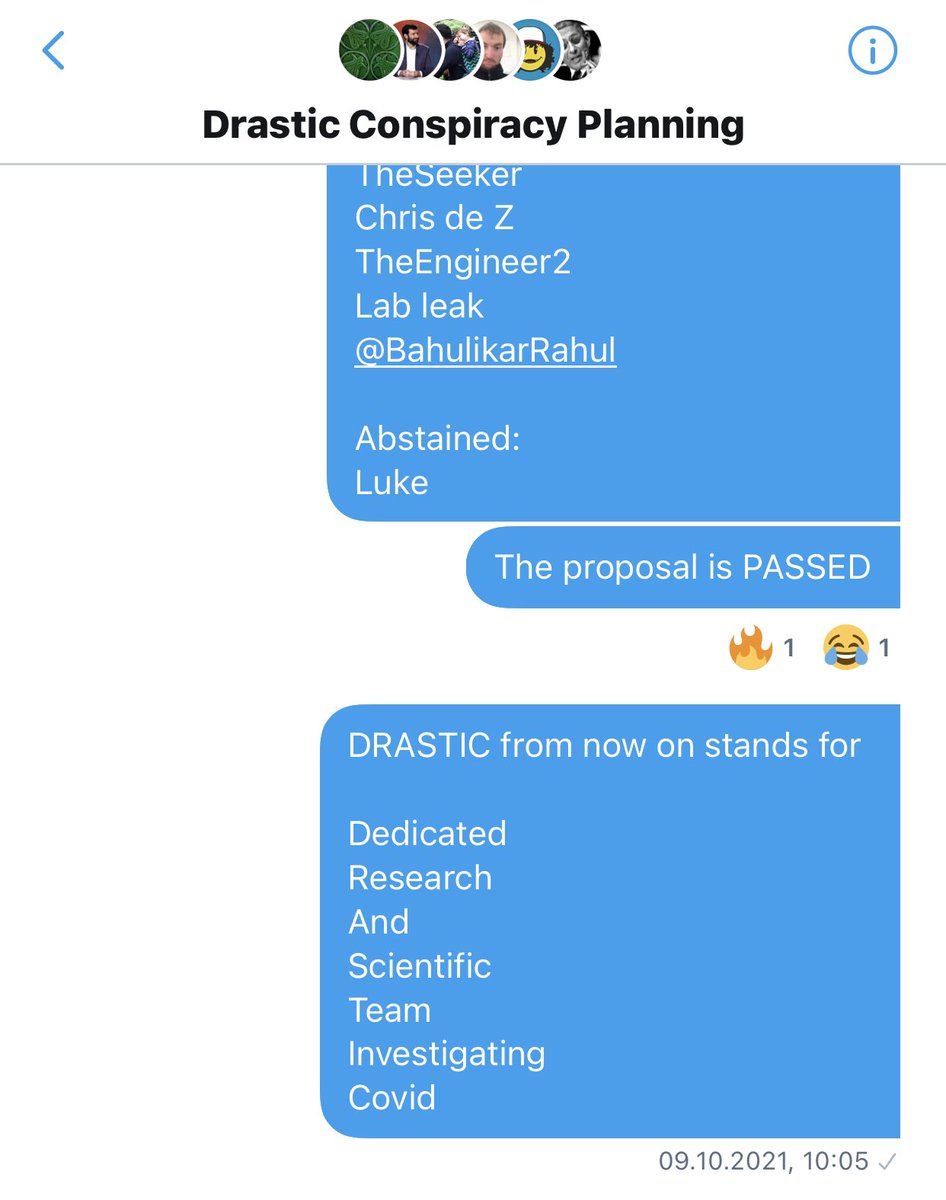
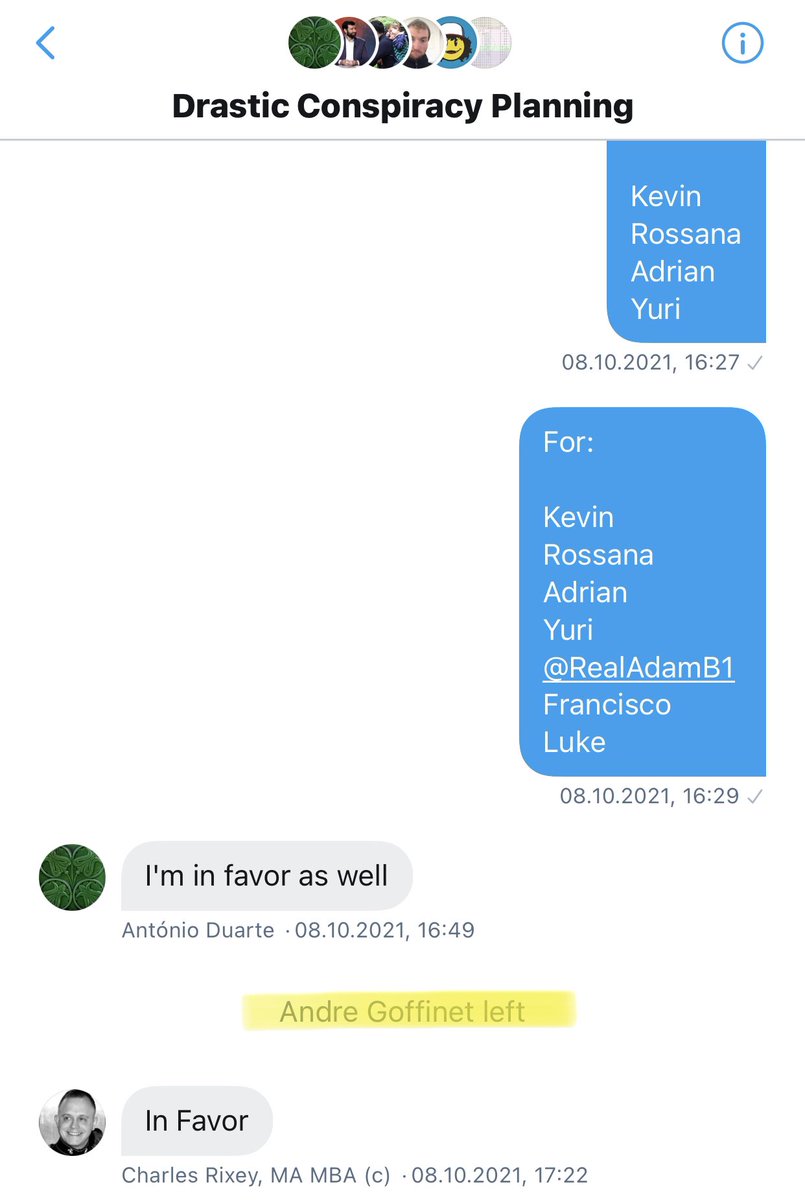
Changing what DRASTIC stands for was proposed *after* the decision to split off from Billy, and was put to a vote in the original DRASTIC group, to which we were added back once Billy was removed. https://t.co/lksMdfRVUm
@ydeigin - Yuri Deigin
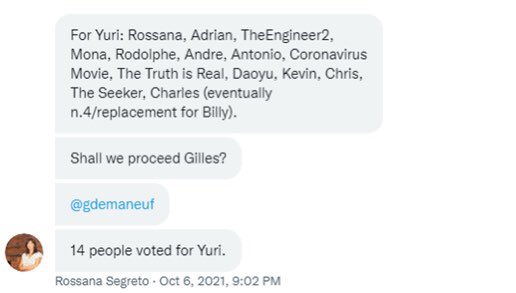
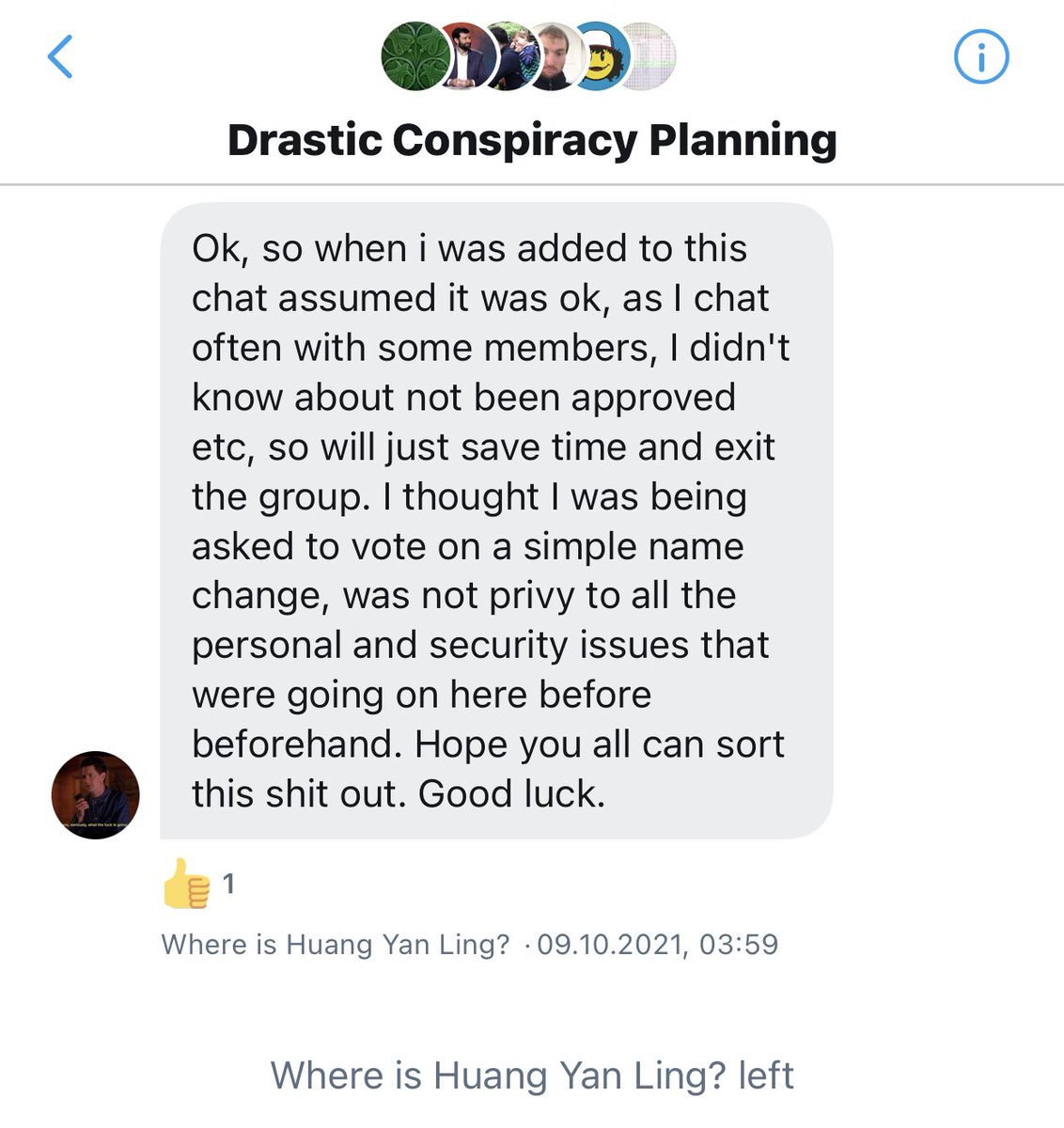
As Andre and WhereIsHuangYanling left the group before that vote, 21 members remained, so 13 votes for and 1 abstention were enough to pass the proposal. https://t.co/P2CovhOwEx
@ydeigin - Yuri Deigin
In closing, we never wanted to make this dirty laundry public, but with Billy’s misinformation campaign trying to portray the split as a result of disagreement on the name change we had to set the record straight. And no, rebranding was not my idea, although I fully support it.
@ydeigin - Yuri Deigin
Also, I am not trying to become “the leader of DRASTIC” or portray myself as some sort of “coordinator” — DRASTIC is and will remain decentralized and fully democratic. But I *will* always stand up to bullying, as that is something I simply cannot tolerate.

@ydeigin - Yuri Deigin
4 years after my first Medium article on Covid origins, I wrote another one. In it, I make the case that SARS-CoV-2 is precisely the virus WIV was hunting for in 2019. https://yurideigin.medium.com/sars-cov-2-is-precisely-the-virus-wiv-was-hunting-for-in-2019-b93b6c996136 Below is a thread on the key points:


@ydeigin - Yuri Deigin
4/ Now, the loss of the N370 glycan by SARS2 has been shown to greatly increase its infectivity in human cells: “Using a reverse genetics system to generate a SARS-CoV-2 mutant containing the putative ancestral SNP, we show that the A372T S mutant virus replicates over 60-fold less efficiently than WT SARS-CoV-2 in Calu-3 human lung epithelial cells (Figure 4d). Further, growth of the A372T S mutant was reduced greatly for multiple days, which may be indicative of an effect on viral shedding kinetics in humans. We also generated the D614G S mutant here—reported widely to increase SARS-CoV-2 infectivity (Korber et al., 2020)—which only increased viral titers by a maximum of 2.9-fold in Calu-3 cells compared with the WT, a finding that is consistent with previous results (Plante et al., 2021).” https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8260498/


@ydeigin - Yuri Deigin
3/ Circling back to the DEFUSE proposal, the N370 glycan in SARS2 is the same glycan as N357 in SARS1 which was implicated as being important for DC-SIGN binding in 2006: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1641789/


@ydeigin - Yuri Deigin
2/ What the 2007 paper did not know at the time that the DEFUSE authors pointed out is that the bat progenitor strains like WIV1/Rs3367 or SHC014 also have glycans at those positions. This is what likely made the DEFUSE authors interested in the host jumping potential of these glycans and potentially genetically modifying them to further study their role:

@ydeigin - Yuri Deigin
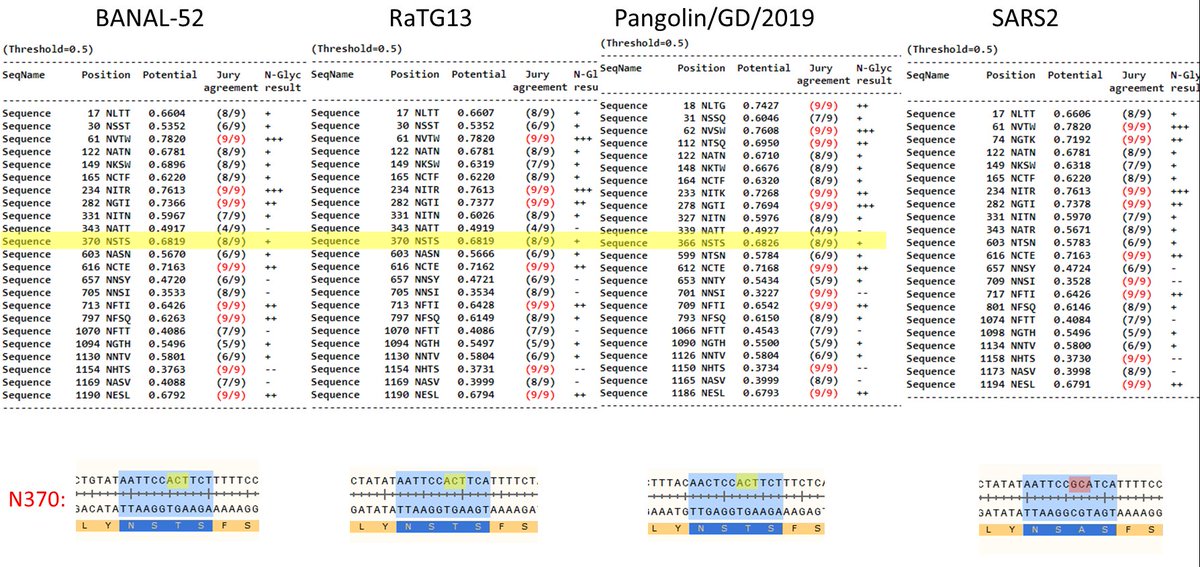
Here are the N-linked glycans in SARS2 vs. related bat or pangolin CoVs: all have the N370 glycan except SARS2. https://t.co/g3D8XHIgDO

@ydeigin - Yuri Deigin
7/end PS: As an aside, the fact that within months of the SARS1 outbreak, 5 different civet SARS1 progenitor strains were identified, but 3.5 years after the SARS2 outbreak we have nothing even remotely close to an intermediate host strain or even a potential bat progenitor — despite an additional two decades of progress in sequencing technology — only keeps adding to my skepticism about a natural origin of SARS2.

@ydeigin - Yuri Deigin
6/ All of this begs the question: how likely is it that these unique features of SARS2 — the FCS and the ablated N370 glycan — unseen in any natural SARS-like virus and unlikely to arise in bats due to selective pressure against them are the result of DEFUSE-inspired genetic engineering? Alternatively, could they arise via serial passaging in civets or their cells? As DEFUSE stated, early SARS1 strains showed a loss of N-linked glycans, and WIV is known to have conducted infectivity experiments on live civets using SARS-like viruses:

@ydeigin - Yuri Deigin
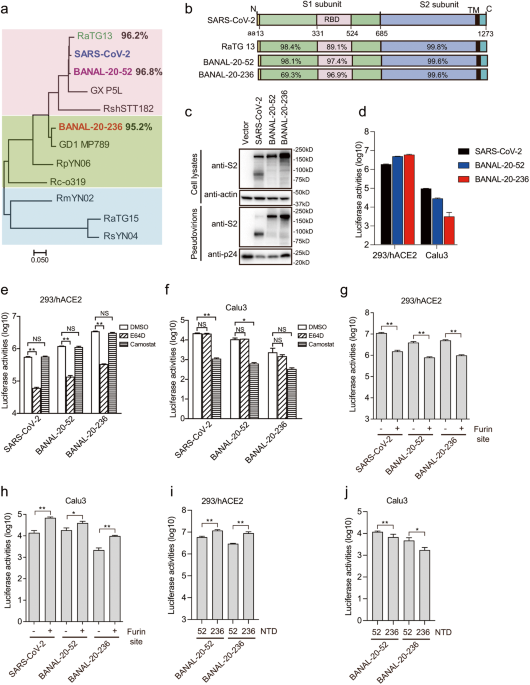
5/ However, this mutation is unlikely to have arisen in bats as it is detrimental to oral-fecal transmission (which SARS-like CoVs rely on in bats; this is also likely why we don’t see an FCS in bat SARS-like CoVs): “Why do all bat SC2r-CoVs retain T372, not A372, in their spike proteins, even though the A372 mutant showed substantially higher infectivity than T372? Since the fecal-oral route plays a vital role in bat CoV transmission among bats31,32, we hypothesized that fecal-oral transmission might favor S proteins in all "down" conformation during natural selection, and T372A change might cause some RBDs to assume “up” conformation, which might be detrimental for the survival of S proteins during their passage through the bat stomach. The pH of an insectivorous bat stomach is around 5.633. To test this hypothesis, WT and T372A mutant S pseudovirions were treated with TPCK trypsin at pH 5.5 at 37 °C, a condition roughly mimicking bat stomach digestion. With increase of trypsin concentration, both WT and T372A pseudovirions lost significant amount of infectivity (Fig. 4b, c). However, the speed and extent of infectivity loss varied significantly between WT and T372A mutants (Fig. 4b, c). While a brief 10 min treatment of trypsin at 2.5 μg/mL resulted in over 96.6% and 99.9% loss of infectivity for BANAL-20-52 T372A and BANAL-20-236 T368A mutants, respectively, WT BANAL-20-52 and BANAL-20-236 S pseudovirions retained more than 37% and 21% of infectivity (Fig. 4b, c). Moreover, even after 40 min digestion with trypsin at 2.5 μg/mL, WT BANAL-20-52 and BANAL-20-236 pseudoviruses still retain over 23% and 14% of infectivity, respectively, whereas T372A and T368A mutants almost completely lost infectivity (Fig. 4d, e).” https://www.nature.com/articles/s41421-023-00581-9

@ydeigin - Yuri Deigin
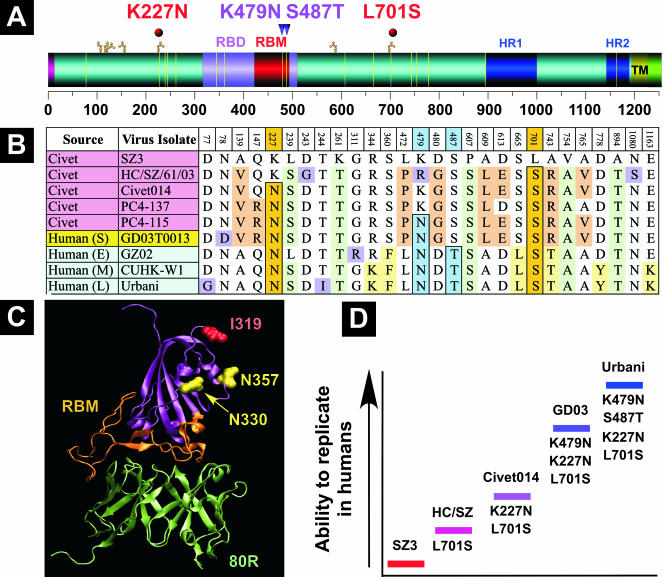
1/ Besides the furin cleavage site (FCS), SARS2 has another unique feature mentioned in DEFUSE not yet seen in any natural SARS-like viruses – an ablated N-linked glycan at position N370. This glycan was ablated via a T372A amino acid mutation that came about via a double nucleotide mutation of the original ACT codon into GCA (the latter, incidentally, is the same codon as the one coding for alanine – out of 4 possible alanine codons – in the PRRA insertion which has created an FCS in SARS2). Importantly, the T327A mutation greatly increases SARS2 infectivity in human lung cells but, just like an FCS, this kind of a mutation seems to have selective pressure AGAINST it in ancestral bat viruses. DEFUSE’s interest in N-linked glycans stems from a very curious observation about SARS1 whose bat progenitor seems to have temporarily lost two of its N-linked glycans in civet SARS1 progenitors before re-acquiring them, and this led virologists to hypothesize that those glycans could be relevant for host switching. This is described in DEFUSE in a somewhat convoluted way: “N-linked glycosylation: Some glycosylation events regulate SARS-CoV particle binding DC-SIGN/L-SIGN, alternative receptors for SARS-CoV entry into macrophages or monocytes [76,77]. Mutations that introduced two new N-linked glycosylation sites may have been involved in the emergence of human SARS-CoV from civet and raccoon dogs [77]. While the sites are absent from civet and raccoon dog strains and clade 2 SARSr-CoV, they are present in WIV1, WIV16 and SHC014, supporting a potential role for these sites in host jumping. To evaluate this, we will sequentially introduce clade 2 disrupting residues of SARS-CoV and SHC014 and evaluate virus growth in Vero cells, nonpermissive cells ectopically expressing DC-SIGN, and in human monocytes and macrophages anticipating reduced virus growth efficiency. We will introduce the clade I mutations that result in N-linked glycosylation in rs4237 RBD deletion repaired strains, evaluating virus growth efficiency in HAE, Vero cells, or nonpermissive cells +/- ectopic DC-SIGN expression [77]. In vivo, we will evaluate pathogenesis in transgenic hACE2 mice.” The [77] paper cited in DEFUSE is a 2007 work by Han et al. titled “Specific Asparagine-Linked Glycosylation Sites Are Critical for DC-SIGN- and L-SIGN-Mediated Severe Acute Respiratory Syndrome Coronavirus Entry”. It looked at the 5 civet progenitor strains of SARS1 and showed that initially those strains did not have glycans around positions N227 and N699 but then eventually acquired them in civet progenitors and kept in human SARS1. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2168787/


@ydeigin - Yuri Deigin
I was quite shocked to discover that Yaroslav Hunka — the Nazi veteran invited to the House of Commons — was actually previously honored (along with several dozen other Waffen SS veterans) by the Ukrainian Canadian Congress in 2007. Notably, the Ukrainian ambassador to Canada was a special guest at the ceremony and gave a celebratory speech: https://diasporiana.org.ua/wp-content/uploads/books/13151/file.pdf